Difference between revisions of "Davidson Missouri W"
Amshoecraft (talk | contribs) (→Our Project) |
|||
| (70 intermediate revisions by 3 users not shown) | |||
| Line 1: | Line 1: | ||
| − | [[ | + | <center>[[Davidson Missouri W| <span style="color:black">Home</span>]] | [[Davidson Missouri W/Background Information| <span style="color:red">Background Information</span>]] | [[Davidson Missouri W/Solving the HPP in vivo| <span style="color:red">Current Project: Solving the Hamiltonian Path Problem ''in vivo''</span>]] | [[Davidson Missouri W/Mathematical Modeling| <span style="color:red">Mathematical Modeling</span>]] | [[Davidson Missouri W/Gene splitting| <span style="color:red"> Gene Splitting </span>]] | [[Davidson Missouri W/Controlling Expression| <span style="color:red"> Controlling Expression </span>]] | [[Davidson Missouri W/Traveling Salesperson Problem| <span style="color:red">Traveling Salesperson Problem</span> ]] | [[Davidson Missouri W/Backwards promotion and read-through transcription| <span style="color:red">Backwards Promotion and Read-Through Transcription</span>]] | [[Davidson Missouri W/Software|<span style="color:red">Software</span>]] | [[Davidson Missouri W/Resources and Citations|<span style="color:red">Resources and Citations</span>]]</center> |
| − | {| border="1" cellpadding="5" cellspacing="0" align="center" width=" | + | <hr> |
| − | + | <br> | |
| + | |||
| + | [[Image:DMW-1.jpg|300px|center]] | ||
| + | |||
| + | <center> | ||
| + | =The Team= | ||
| + | </center> | ||
| + | {| border="1" cellpadding="5" cellspacing="0" align="center" width="100%" | ||
|- | |- | ||
| − | ! style="color: | + | ! style="color: white; background-color: black;"| The Team |
| − | ! | + | ! style="color: white; background-color: black;" | The Faculty |
| + | ! style="color: white; background-color: black;" | Team Logos | ||
| + | ! style="color: white; background-color: black;" | Group Photo | ||
|- | |- | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | + | |style="color: black; background-color: red;" align="center"| '''Davidson''' | |
| + | <b> | ||
| + | [[Davidson Missouri W/Oyinade Adefuye|<span style="color:black">Oyinade Adefuye</span>]] | ||
| + | <br> | ||
| + | [[Davidson Missouri W/Will DeLoache|<span style="color:black">Will DeLoache</span>]] | ||
| + | <br> | ||
| + | [[Davidson Missouri W/Jim Dickson|<span style="color:black">Jim Dickson</span>]] | ||
| + | <br> | ||
| + | [[Davidson Missouri W/Andrew Martens|<span style="color:black">Andrew Martens</span>]] | ||
| + | <br> | ||
| + | [[Davidson Missouri W/Amber Shoecraft|<span style="color:black">Amber Shoecraft</span>]] | ||
| + | <br> | ||
| + | [[Davidson Missouri W/Mike Waters|<span style="color:black">Mike Waters</span>]] | ||
| + | </b> | ||
| − | [[ | + | |style="color: black; background-color: red;" align="center"| |
| + | <b> | ||
| + | [[Davidson Missouri W/A. Malcolm Campbell|<span style="color:black">A. Malcom Campbell</span>]] | ||
| + | <br> | ||
| + | [[Davidson Missouri W/Karmella Haynes|<span style="color:black">Karmella Haynes</span>]] | ||
<br> | <br> | ||
| − | [[ | + | [[Davidson Missouri W/Laurie Heyer|<span style="color:black">Laurie Heyer</span>]] |
| − | + | </b> | |
| − | < | + | |
| − | + | |style="color: black; background-color: white;" align="center"| | |
| − | | | + | [[Image:DavidsonLogo.gif]] |
| + | |||
| + | |style="color: black; background-color: white;" align="center"| | ||
| + | [[Image:Team1.jpg|thumb|300px]] | ||
| + | |||
|- | |- | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | [ | + | |style="color: black; background-color: gold;" align="center"|'''Missouri Western''' |
| + | <b> | ||
| + | [[Davidson Missouri W/Jordan Baumgardner|<span style="color:black;">Jordan Baumgardner</span>]] | ||
| + | <br> | ||
| + | [[Davidson Missouri W/Tom Crowley|<span style="color:black;">Tom Crowley</span>]] | ||
| + | <br> | ||
| + | [[Davidson Missouri W/Lane H. Heard|<span style="color:black;">Lane H. Heard</span>]] | ||
| + | <br> | ||
| + | [[Davidson Missouri W/Nickolaus Morton|<span style="color:black;">Nickolaus Morton</span>]] | ||
| + | <br> | ||
| + | [[Davidson Missouri W/Michelle Ritter|<span style="color:black;">Michelle Ritter</span>]] | ||
| + | <br> | ||
| + | [[Davidson Missouri W/Jessica Treece|<span style="color:black;">Jessica Treece</span>]] | ||
| + | <br> | ||
| + | [[Davidson Missouri W/Matthew Unzicker|<span style="color:black;">Matthew Unzicker</span>]] | ||
| + | <br> | ||
| + | [[Davidson Missouri W/Amanda Valencia|<span style="color:black;">Amanda Valencia</span>]] | ||
| + | </b> | ||
| + | |||
| + | |style="color: black; background-color: gold;" align="center"| | ||
| + | <b> | ||
| + | [[Davidson Missouri W/Todd Eckdahl|<span style="color:black;">Todd Eckdahl</span>]] | ||
| + | <br> | ||
| + | [[Davidson Missouri W/Jeff Poet|<span style="color:black;">Jeff Poet</span>]] | ||
| + | </b> | ||
| − | [ | + | |style="color: black; background-color: white;" align="center"|[[Image:MWLogo.gif]] |
| − | + | |style="color: black; background-color: white;" align="center"| | |
| + | |- | ||
| − | + | |} | |
| − | + | <br> | |
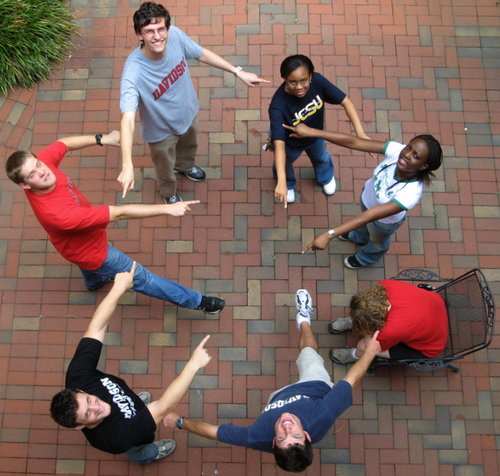
| − | + | [[Image:Team_graph.jpg|center|thumb|500px|A Human Representation of Adleman's Graph (see below)]] | |
| − | + | <br><br> | |
| − | |||
| − | + | <center> | |
| − | |||
| − | + | =Our Project= | |
| − | + | </center> | |
| − | + | {| border="1" cellpadding="5" cellspacing="0" align="center" width="90%" | |
| − | [ | + | |- |
| + | ! style="color: black; background-color: red;" width="20%"| <font size="+1">In Depth</font> | ||
| + | ! colspan="3" style="color: black; background-color: red;" width="60%"| <font size="+1">Overview</font> | ||
| + | |- | ||
| + | |style="color: black; background-color: black;" align="center"| | ||
| + | [[Davidson Missouri W/Background Information|<span style="color:red">Background Information</span>]] | ||
| + | <br><br><br> | ||
| + | [[Davidson Missouri W/Solving the HPP in vivo|<span style="color:red">Current Project: Solving the Hamiltonian Path Problem ''in vivo''</span>]] | ||
| + | <br><br><br> | ||
| + | [[Davidson Missouri W/Mathematical Modeling|<span style="color:red">Mathematical Modeling</span>]] | ||
| + | <br><br><br> | ||
| + | [[Davidson Missouri W/Gene splitting|<span style="color:red">Gene Splitting</span>]] | ||
| + | <br><br><br> | ||
| + | [[Davidson Missouri W/Controlling Expression|<span style="color:red">Controlling Expression</span>]] | ||
| + | <br><br><br> | ||
| + | [[Davidson Missouri W/Traveling Salesperson Problem|<span style="color:red">Traveling Salesperson Problem</span>]] | ||
| + | <br><br><br> | ||
| + | [[Davidson Missouri W/Backwards Promotion and read-through transcription|<span style="color:red">Backwards Promotion and Read-Through Transcription</span>]] | ||
| + | <br><br><br> | ||
| + | [[Davidson Missouri W/Resources and Citations|<span style="color:red">Resources and Citations</span>]] | ||
| + | <br><br><Br> | ||
| + | |Hamiltonian Path Problem | ||
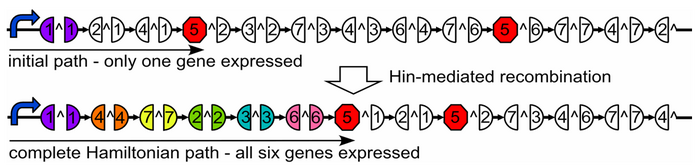
| + | As a part of iGEM2006, a combined team from Davidson College and Missouri Western State University reconstituted a hin/hix DNA recombination mechanism which exists in nature in Salmonella as standard biobricks for use in ''E. coli''. The purpose of the 2006 combined team was to provide a proof of concept for a bacterial computer in using this mechanism to solve a variation of The Pancake Problem from Computer Science. This task utilized both biology and mathematics students and faculty from the two institutions. | ||
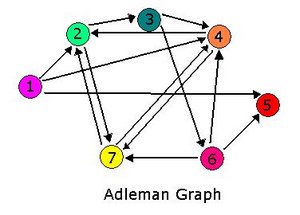
| − | + | For 2007, we continue our collaboration and our efforts to manipulate ''E. coli'' into mathematics problem solvers as we refine our efforts with the hin/hix mechanism to explore another mathematics problem, the Hamiltonian Path Problem. This problem was the subject of a groundbreaking paper by Adelman in 1994 (citation below) where a unique Hamiltonian path was found in vitro for a particular directed graph on seven nodes. We propose to make progress toward solving the particular problem in vivo. | |
| − | |||
| − | + | <br> | |
| − | + | [[Image:AdelmanGraph.JPG|thumb|300px|center|The Adleman graph.]] | |
| + | <br> | ||
| + | [[Image:HamiltonianGraph.PNG|thumb|700px|center|A graph implemented on a plasmid.]] | ||
| − | + | |} | |
Latest revision as of 15:22, 9 August 2007
The Team
| The Team | The Faculty | Team Logos | Group Photo |
|---|---|---|---|
| Davidson
Oyinade Adefuye
|
Error creating thumbnail: Unable to save thumbnail to destination
|
||
| Missouri Western
Jordan Baumgardner
|
Error creating thumbnail: Unable to save thumbnail to destination
|
Our Project
| In Depth | Overview | ||
|---|---|---|---|
|
Background Information
|
Hamiltonian Path Problem
As a part of iGEM2006, a combined team from Davidson College and Missouri Western State University reconstituted a hin/hix DNA recombination mechanism which exists in nature in Salmonella as standard biobricks for use in E. coli. The purpose of the 2006 combined team was to provide a proof of concept for a bacterial computer in using this mechanism to solve a variation of The Pancake Problem from Computer Science. This task utilized both biology and mathematics students and faculty from the two institutions. For 2007, we continue our collaboration and our efforts to manipulate E. coli into mathematics problem solvers as we refine our efforts with the hin/hix mechanism to explore another mathematics problem, the Hamiltonian Path Problem. This problem was the subject of a groundbreaking paper by Adelman in 1994 (citation below) where a unique Hamiltonian path was found in vitro for a particular directed graph on seven nodes. We propose to make progress toward solving the particular problem in vivo.
| ||