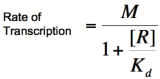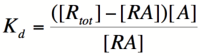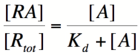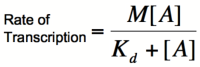Difference between revisions of "CellularMemory:Mathematical Models"
Wideloache (talk | contribs) |
Wideloache (talk | contribs) |
||
| Line 10: | Line 10: | ||
When dealing with gene activation and repression networks, such as those described in the common [[CellularMemory:Biological Designs |Biological Designs]] for synthetic cellular memory section of this paper, mathematical models are mainly used to model the transcription rate of a gene(s). Three common models that describe transcription rate as a function of repressor or activator are, in order of increasing complexity, the [http://en.wikipedia.org/wiki/Michaelis-Menten_kinetics Michaelis-Menten] model, the [http://en.wikipedia.org/wiki/Hill_equation Hill] model, and the [http://en.wikipedia.org/wiki/MWC_model Monod-Wymann-Changeux] model. Each of these model builds on the one preceding it to paint a clearer picture of how binding occurs between two or more molecules and how that binding affects transcription rates. | When dealing with gene activation and repression networks, such as those described in the common [[CellularMemory:Biological Designs |Biological Designs]] for synthetic cellular memory section of this paper, mathematical models are mainly used to model the transcription rate of a gene(s). Three common models that describe transcription rate as a function of repressor or activator are, in order of increasing complexity, the [http://en.wikipedia.org/wiki/Michaelis-Menten_kinetics Michaelis-Menten] model, the [http://en.wikipedia.org/wiki/Hill_equation Hill] model, and the [http://en.wikipedia.org/wiki/MWC_model Monod-Wymann-Changeux] model. Each of these model builds on the one preceding it to paint a clearer picture of how binding occurs between two or more molecules and how that binding affects transcription rates. | ||
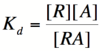
| − | The Hill equation is used most often to model cellular memory networks. In order to describe how the Hill equation works, it is first helpful to examine a simpler model. Based on the two [[CellularMemory:Biological Designs |biological designs]] presented in the previous section, it can be reasoned that to mathematically model these designs, we must be able to describe the effect of repressor/promoter binding on transcription rates ([[CellularMemory:Biological Designs#Mutual Repression |mutual repression]]) and the effect of activator/repressor binding on transcription rates ([[CellularMemory:Biological Designs#Autoregulatory Positive Feedback |autoregulatory positive feedback]]). | + | The Hill equation is used most often to model cellular memory networks. In order to describe how the Hill equation works, it is first helpful to examine a simpler model. Based on the two [[CellularMemory:Biological Designs |biological designs]] presented in the previous section, it can be reasoned that to mathematically model these designs, we must be able to describe the effect of repressor/promoter binding on transcription rates ([[CellularMemory:Biological Designs#Mutual Repression |mutual repression]]) and the effect of activator/repressor binding on transcription rates ([[CellularMemory:Biological Designs#Autoregulatory Positive Feedback |autoregulatory positive feedback]]). The two binding equations presented below do just that. |
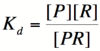
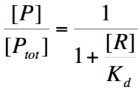
===Repressor/Promoter Binding=== | ===Repressor/Promoter Binding=== | ||
Revision as of 23:05, 29 November 2007
Main Page | Biological Designs | Mathematical Models | Toggle Switch | Hysteresis | Permanent Memory | Conclusions | References
Contents
|