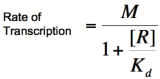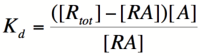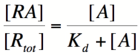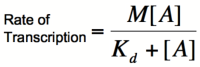Difference between revisions of "CellularMemory:Mathematical Models"
Wideloache (talk | contribs) (→Determining the Optimal Values of Other Parameters) |
Wideloache (talk | contribs) |
||
| Line 5: | Line 5: | ||
Mathematical modeling is an important component in the construction of rationally designed gene networks. The complexity of biological systems makes necessary the use of sophisticated mathematical models to accurately predict network functionality. Models are useful in tuning the individual components of a network to increase system robustness. They also provide a basis for comparison of experimental results with expected results, allowing researchers to test the validity of their assumptions. | Mathematical modeling is an important component in the construction of rationally designed gene networks. The complexity of biological systems makes necessary the use of sophisticated mathematical models to accurately predict network functionality. Models are useful in tuning the individual components of a network to increase system robustness. They also provide a basis for comparison of experimental results with expected results, allowing researchers to test the validity of their assumptions. | ||
| − | + | =Modeling Transcription Rates= | |
Note: the following mathematical analysis was developed based on a summary of the input function of a gene in ''An Introduction to Systems Biology: Design Principles of Biological Circuits'' [[CellularMemory:References |(Alon, pgs. 241-250)]]. | Note: the following mathematical analysis was developed based on a summary of the input function of a gene in ''An Introduction to Systems Biology: Design Principles of Biological Circuits'' [[CellularMemory:References |(Alon, pgs. 241-250)]]. | ||
| Line 12: | Line 12: | ||
The Hill equation is used most often to model cellular memory networks. In order to describe how the Hill equation works, it is first helpful to examine a simpler model. Based on the two [[CellularMemory:Biological Designs |biological designs]] presented in the previous section, it can be reasoned that to mathematically model these designs, we must be able to describe the effect of repressor/promoter binding on transcription rates (for [[CellularMemory:Biological Designs#Mutual Repression |mutual repression]] designs) and the effect of activator/repressor binding on transcription rates (for [[CellularMemory:Biological Designs#Autoregulatory Positive Feedback |autoregulatory positive feedback]] designs). The two binding equations presented below do just that. | The Hill equation is used most often to model cellular memory networks. In order to describe how the Hill equation works, it is first helpful to examine a simpler model. Based on the two [[CellularMemory:Biological Designs |biological designs]] presented in the previous section, it can be reasoned that to mathematically model these designs, we must be able to describe the effect of repressor/promoter binding on transcription rates (for [[CellularMemory:Biological Designs#Mutual Repression |mutual repression]] designs) and the effect of activator/repressor binding on transcription rates (for [[CellularMemory:Biological Designs#Autoregulatory Positive Feedback |autoregulatory positive feedback]] designs). The two binding equations presented below do just that. | ||
| − | + | ==Repressor/Promoter Binding== | |
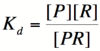
Let's start by looking at the effect of repressor/promoter binding on transcription rate. The starting point for this proof is the [http://en.wikipedia.org/wiki/Dissociation_constant dissociation constant] for promoter and repressor binding. This constant, K<sub>d</sub>, measures the tendency for a repressor/promoter complex to fall apart into its two separate subunits. The value of K<sub>d</sub> is determined by the binding affinity of the two molecules and is written as: | Let's start by looking at the effect of repressor/promoter binding on transcription rate. The starting point for this proof is the [http://en.wikipedia.org/wiki/Dissociation_constant dissociation constant] for promoter and repressor binding. This constant, K<sub>d</sub>, measures the tendency for a repressor/promoter complex to fall apart into its two separate subunits. The value of K<sub>d</sub> is determined by the binding affinity of the two molecules and is written as: | ||
| Line 33: | Line 33: | ||
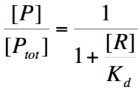
Based on the equation above, it can be seen that the dissociation constant, K<sub>d</sub>, equals the repressor concentration at which the transcription rate is half of its maximal value. The value of K<sub>d</sub> can, therefore, easily be determined experimentally. | Based on the equation above, it can be seen that the dissociation constant, K<sub>d</sub>, equals the repressor concentration at which the transcription rate is half of its maximal value. The value of K<sub>d</sub> can, therefore, easily be determined experimentally. | ||
| − | + | ==Activator/Repressor Binding (The Michaelis-Menten Equation)== | |
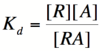
In order to describe transcription rates of an autoregulatory positive feedback system, we must look at the effect of activator/repressor binding on promoter activity. This proof works much in the same way as the repressor/promoter binding equation and it will lead us in the end to the Michaelis-Menten equation. We will start with the dissociation constant, K<sub>d</sub>, for activator/repressor binding: | In order to describe transcription rates of an autoregulatory positive feedback system, we must look at the effect of activator/repressor binding on promoter activity. This proof works much in the same way as the repressor/promoter binding equation and it will lead us in the end to the Michaelis-Menten equation. We will start with the dissociation constant, K<sub>d</sub>, for activator/repressor binding: | ||
| Line 54: | Line 54: | ||
This equation is known as the Michaelis-Menten equation. Once again, it is true that the dissociation constant, K<sub>d</sub>, equals the activator concentration at which the transcription rate is half of its maximal value. The value of K<sub>d</sub> can, therefore, easily be determined experimentally. | This equation is known as the Michaelis-Menten equation. Once again, it is true that the dissociation constant, K<sub>d</sub>, equals the activator concentration at which the transcription rate is half of its maximal value. The value of K<sub>d</sub> can, therefore, easily be determined experimentally. | ||
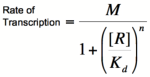
| − | + | ==The Hill Equation== | |
For both of the equations derived above, it can be reasoned that when high levels of repressor or activator are in their respective systems, the rate of change in transcription rate is relatively low. This is the result that we would expect, because at high levels of repressor or activator, the systems would become saturated and would be minimally affected by the addition of more repressor or activator. Unfortunately, the above equations do not accurately depict many systems when repressor or activator concentrations are low. This is because of the tendency of transcription factors to be composed of multiple subunits. While a single subunit (a monomer) can bind to its target molecule by itself, in order to achieve maximal binding affinity, it is often the case that multiple subunits (dimers, tetramers, etc.) are required. This phenomenon is referred to as the cooperativity of binding because it refers to multiple activator or repressor subunits working together to bind as tightly as possible. The equations above do not take cooperativity into account, however, the Hill equation modifies them so that cooperativity can be included in the description of binding. | For both of the equations derived above, it can be reasoned that when high levels of repressor or activator are in their respective systems, the rate of change in transcription rate is relatively low. This is the result that we would expect, because at high levels of repressor or activator, the systems would become saturated and would be minimally affected by the addition of more repressor or activator. Unfortunately, the above equations do not accurately depict many systems when repressor or activator concentrations are low. This is because of the tendency of transcription factors to be composed of multiple subunits. While a single subunit (a monomer) can bind to its target molecule by itself, in order to achieve maximal binding affinity, it is often the case that multiple subunits (dimers, tetramers, etc.) are required. This phenomenon is referred to as the cooperativity of binding because it refers to multiple activator or repressor subunits working together to bind as tightly as possible. The equations above do not take cooperativity into account, however, the Hill equation modifies them so that cooperativity can be included in the description of binding. | ||
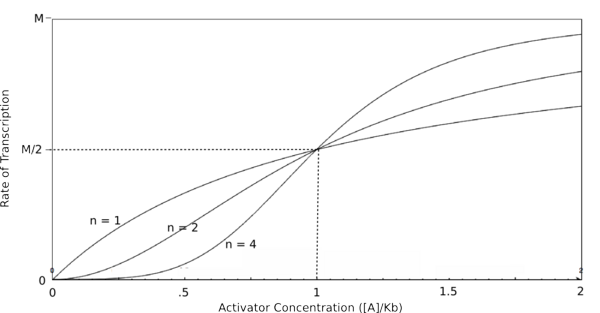
| Line 67: | Line 67: | ||
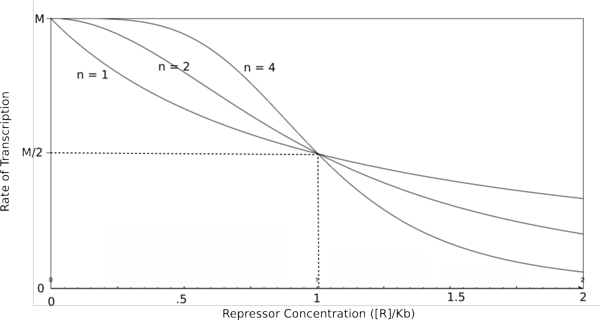
In the graphs above, three things should be noted. First, when K<sub>b</sub> is equal to [R] or [A] (meaning that [R]/K<sub>b</sub> or [A]/K<sub>b</sub> equals 1), the rate of transcription is equal to half of the maximal value (M/2). This trend is denoted by the dotted line. Second, graphs of activators and repressors with a Hill coefficient of 1 are described by the Michaelis-Menten model (not taking cooperativity into account). Therefore, the Hill equation curves for n = 1 also represent the curves for the two relatively simple equations derived earlier in this section. Third, as n increases, the curves develop more and more [http://en.wikipedia.org/wiki/Sigmoid_function sigmoidal] (S-shaped) character, where the curves converge to two different values asymptotically. In the next section, this sigmoidal shape will be shown to be crucial in the establishment of bistability in a memory network. | In the graphs above, three things should be noted. First, when K<sub>b</sub> is equal to [R] or [A] (meaning that [R]/K<sub>b</sub> or [A]/K<sub>b</sub> equals 1), the rate of transcription is equal to half of the maximal value (M/2). This trend is denoted by the dotted line. Second, graphs of activators and repressors with a Hill coefficient of 1 are described by the Michaelis-Menten model (not taking cooperativity into account). Therefore, the Hill equation curves for n = 1 also represent the curves for the two relatively simple equations derived earlier in this section. Third, as n increases, the curves develop more and more [http://en.wikipedia.org/wiki/Sigmoid_function sigmoidal] (S-shaped) character, where the curves converge to two different values asymptotically. In the next section, this sigmoidal shape will be shown to be crucial in the establishment of bistability in a memory network. | ||
| − | + | ==The Monod-Wymann-Changeux Equation== | |
It should be noted briefly that, while the Hill equation does account for cooperativity, it is not always the most accurate description of these types of biological systems. The [http://en.wikipedia.org/wiki/MWC_model Monod-Wymann-Changeux] model improves on the Hill equation by taking into account a decrease in the dissociation constant as more repressor or activator subunits are bound onto a given complex. This model, however, is not frequently used in modeling synthetic cellular memory networks, where the Hill equation appears to suffice. | It should be noted briefly that, while the Hill equation does account for cooperativity, it is not always the most accurate description of these types of biological systems. The [http://en.wikipedia.org/wiki/MWC_model Monod-Wymann-Changeux] model improves on the Hill equation by taking into account a decrease in the dissociation constant as more repressor or activator subunits are bound onto a given complex. This model, however, is not frequently used in modeling synthetic cellular memory networks, where the Hill equation appears to suffice. | ||
| + | |||
| + | |||
| + | =Determining the Optimal Values of Parameters= | ||
| + | The previous section presented mathematical models that are commonly used to describe how transcription rates depend on the concentration of repressors and activators. Now that we understand the origin of these individual equations, we can more easily understand how they are integrated into a system of equations that describes a larger gene network. The power of modeling comes in its ability to determine what values are needed for different system parameters in order to produce a working system. Because these systems of equations must be specific to the biological design, I will not describe them in detail. However, understanding how the above equations were derived will provide us with the tools to understand the models behind many different biological designs. Below, I will present one example of a system of equations that describes the construction of bistability in a synthetic cellular memory network. This system of equations will show that cooperativity of binding is necessary for the construction of bistability for an autoregulatory positive feedback design. In addition to providing information about the necessity of cooperativity, mathematical models also help in determining optimal values for ribosomal binding site (RBS) strength, maximal promoter activity levels, decay rates, and many other parameters. While the derivation of these values will not be discussed explicitly in this paper, know that obtainment of these values is crucial to the rational design of functioning biological networks. | ||
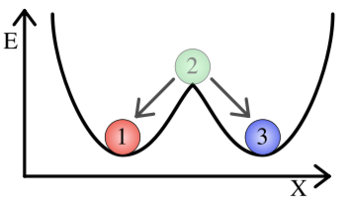
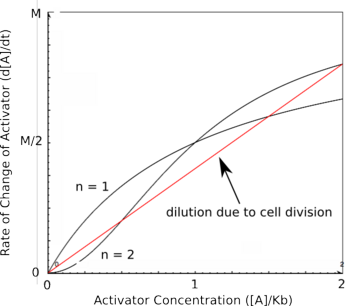
==Cooperativity and Bistability== | ==Cooperativity and Bistability== | ||
| Line 83: | Line 87: | ||
[[Image:linebreak.png]] | [[Image:linebreak.png]] | ||
| − | |||
| − | |||
<hr> | <hr> | ||
Revision as of 19:00, 30 November 2007
Main Page | Biological Designs | Mathematical Models | Toggle Switch | Hysteresis | Permanent Memory | Conclusions | References
|