Difference between revisions of "CellularMemory:Biological Designs"
Wideloache (talk | contribs) (→Overview) |
Wideloache (talk | contribs) (→Autoregulatory Positive Feedback) |
||
| Line 25: | Line 25: | ||
The autoregulatory positive feedback design refers to a gene network that contains a constitutively repressed promoter upstream of its activator gene. This genetic construct creates a positive feedback loop, where initial activation of the promoter causes transcription of an activator gene and thus further increases promoter strength. This positive feedback eventually converges to a steady state of gene expression ([[CellularMemory:Mathematical Models |see mathematical modeling]]). Autoregulatory positive feedback is the biological design used in the final two papers that will be discussed ([[CellularMemory:References |Kramer, 2005]] and [[CellularMemory:References |Ajo-Franklin, 2007]]). | The autoregulatory positive feedback design refers to a gene network that contains a constitutively repressed promoter upstream of its activator gene. This genetic construct creates a positive feedback loop, where initial activation of the promoter causes transcription of an activator gene and thus further increases promoter strength. This positive feedback eventually converges to a steady state of gene expression ([[CellularMemory:Mathematical Models |see mathematical modeling]]). Autoregulatory positive feedback is the biological design used in the final two papers that will be discussed ([[CellularMemory:References |Kramer, 2005]] and [[CellularMemory:References |Ajo-Franklin, 2007]]). | ||
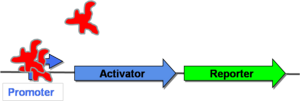
| − | In [http://gcat.davidson.edu/GcatWiki/index.php/Image:Posfeedoff.png Figure 3] to the right, the promoter is initially repressed by a protein that is native to the cell. This means that neither the activator | + | In [http://gcat.davidson.edu/GcatWiki/index.php/Image:Posfeedoff.png Figure 3] to the right, the promoter is initially repressed by a protein that is native to the cell. This means that neither the activator nor the reporter gene will be transcribed. This cell is, therefore, in the "off" state. |
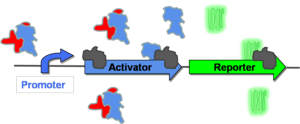
In order to switch to the "on" state ([http://gcat.davidson.edu/GcatWiki/index.php/Image:Posfeedon.png Figure 4]), some stimulus must first allow for some low level of promoter activation. Once the promoter is activated, it will transcribe both its activator gene and the reporter gene. As more activator is produced, higher levels of gene expression will be observed. Eventually, the cell will arrive at a maximum level of gene expression at which point it is considered to be in the "on" state (in this case, green fluorescence). | In order to switch to the "on" state ([http://gcat.davidson.edu/GcatWiki/index.php/Image:Posfeedon.png Figure 4]), some stimulus must first allow for some low level of promoter activation. Once the promoter is activated, it will transcribe both its activator gene and the reporter gene. As more activator is produced, higher levels of gene expression will be observed. Eventually, the cell will arrive at a maximum level of gene expression at which point it is considered to be in the "on" state (in this case, green fluorescence). | ||
Revision as of 18:42, 6 December 2007
Main Page | Biological Designs | Mathematical Models | Toggle Switch | Hysteresis | Permanent Memory | Conclusions | References
|