|
|
| (289 intermediate revisions by 2 users not shown) |
| Line 1: |
Line 1: |
| | + | Austin Mudd - Spring 2013 |
| | + | <br/>Shortened URL: http://goo.gl/zuTkP |
| | + | |
| | + | |
| | __TOC__ | | __TOC__ |
| | | | |
| − | ===To Do=== | + | ==Introduction to Flowering== |
| | + | ====The Process of Flowering==== |
| | + | *Flowering is the "switch from vegetative growth (the production of stems and leaves) to reproductive growth (the production of flowers)" ([http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0010065 Higgins ''et al.'', 2010]) |
| | + | *The “shoot apical meristem starts to produce flowers instead of leaves” ([http://www.sciencedirect.com/science/article/pii/S0092867410004411 Fornara ''et al.'', 2010]) |
| | + | *Occurs “when conditions for pollination and seed development are optimal and consequently most plants restrict flowering to a specific time of year” ([http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0010065 Higgins ''et al.'', 2010]) |
| | + | *”The genes and molecular mechanisms controlling flowering have been extensively studied in the model dicot <i>Arabidopsis thaliana</i>” ([http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0010065 Higgins ''et al.'', 2010]) |
| | + | *In <i>Arabidopsis thaliana</i>, “180 genes have been implicated in flowering-time control based on isolation of loss-of-function mutations or analysis of transgenic plants ... Strikingly, several genes act more than once and in several tissues during floral induction” ([http://www.sciencedirect.com/science/article/pii/S0092867410004411 Fornara ''et al.'', 2010]) |
| | + | |
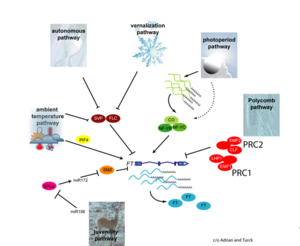
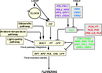
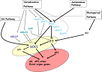
| | + | [[File:Max_Planck_Institute_Pathway.png|thumb|right|alt=Diagram of pathways|The major pathways in the timing of flowering from Turck and Adrian at the [http://www.mpipz.mpg.de/305695/Project_1 Max Planck Institute for Plant Breeding Research]; permission granted]] |
| | + | ====The Timing of Flowering==== |
| | + | *Flowering is controlled by several “major pathways: the photoperiod and vernalization pathways control flowering in response to seasonal changes in day length and temperature; the ambient temperature pathway responds to daily growth temperatures; and the age, autonomous, and gibberellin pathways act more independently of environmental stimuli.” ([http://www.sciencedirect.com/science/article/pii/S0092867410004411 Fornara ''et al.'', 2010]) |
| | + | *These “pathways converge to regulate a small number of ‘floral integrator genes,’ ... which govern flowering time by merging signals from multiple pathways” ([http://www.sciencedirect.com/science/article/pii/S0092867410004411 Fornara ''et al.'', 2010]) |
| | + | |
| | + | ====The Importance of Flowering==== |
| | + | *”Flowering is one of the most important agronomic traits influencing crop yield” ([http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0038250 Jung ''et al.'', 2012]) |
| | + | *”Flowering time is important for adaptation to specific environments and the world's major crop species provide a particularly interesting opportunity for study because they are grown in areas outside the ecogeographical limits of their wild ancestors” ([http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0010065 Higgins ''et al.'', 2010]) |
| | + | *“Adaptation to different environments and practices has been achieved by manipulation of flowering time responses” ([http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0010065 Higgins ''et al.'', 2010]) |
| | + | *The study of flowering is ”critical for the breeding of climate change resilient crop varieties” ([http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0038250 Jung ''et al.'', 2012]) |
| | + | *Flowering is “an excellent system for comparison between and within domestic and wild species” ([http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0010065 Higgins ''et al.'', 2010]) |
| | | | |
| − | Prepare for presentation.
| |
| − | <br/>Aggregate all sequences into a single document in proper FASTA format.
| |
| − | <br/>
| |
| | | | |
| − | ===Chronology of <i>Arabidopsis</i> Flowering Pathways=== | + | ==Pathways Controlling Flowering== |
| | + | ====Age Pathway==== |
| | + | *"The miR156–SPL interaction constitutes an evolutionarily conserved, endogenous cue for both vegetative phase transition and flowering ... The age-dependent decrease in miR156 results in an increase in SPLs that promote juvenile to adult phase transition and flowering through activation of miR172, MADS box genes, and LFY" ([http://www.plantcell.org/content/early/2012/08/29/tpc.112.101014.full.pdf+html Yu ''et al.'', 2012]) |
| | + | *5 <i>Arabidopsis</i> genes are involved in the age pathway: SPL3, SPL4, SPL5, SPL9, SPL10 ([http://onlinelibrary.wiley.com/doi/10.1111/j.1365-313X.2010.04148.x/pdf/ Amasino, 2010]) |
| | | | |
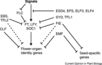
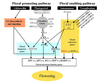
| − | [[Image:Flowering_Pathway_2004.jpeg|700px]]
| + | ====Ambient Temperature Pathway==== |
| − | <br/>Overall pathway for the time of bloom from [http://dev.biologists.org/content/131/16/3829/F1.expansion.html Henderson and Dean, 2004].
| + | *Unlike "the photoperiod and vernalisation pathways [which] monitor seasonal changes in day length or temperature and ... [respond] to exposure to long days or prolonged cold temperatures, the ambient temperature pathway coordinates the response to daily growth temperatures" ([http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0038250 Jung ''et al.'', 2012]) |
| − | <br/><br/> [[Image:Flowering_Pathway_2009.gif|700px]]
| + | *16 <i>Arabidopsis</i> genes are involved in the ambient temperature pathway: AGL31, ATARP6, ATBZIP27, FCA, FD, FLC, FLD, FT, FVE, MAF1, MAF3, MAF4, MAF5, SVP, TFL1, TSF ([http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0038250 Jung ''et al.'', 2012]) |
| − | <br/>Overall pathway for the time of bloom from [http://www.els.net/WileyCDA/ElsArticle/refId-a0002053.html Schneitz, 2009].
| |
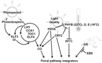
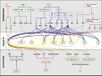
| − | <br/><br/> [[Image:Flowering_Pathway_2010_2011.jpg|700px]]
| |
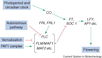
| − | <br/>"The <i>Arabidopsis</i> floral transition network has been delineated as a result of extensive genetic analyses (depicted in a), but genome-wide studies have uncovered new molecular interactions and highlighted the role of AP1 as both a molecular switch and a network hub (shown in b)."
| |
| − | <br/>Overall pathway for the time of bloom from [http://www.sciencedirect.com/science/article/pii/S0168952510001873 Wellmer and Riechmann, 2010] and [http://www.sciencedirect.com/science/article/pii/S0958166910002284 Ferrier et al., 2011].
| |
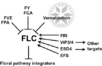
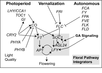
| − | <br/><br/> [[Image:Flowering_Pathway_2012.jpg|700px]]
| |
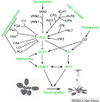
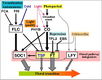
| − | <br/>"Soybean genes marked with * are those that share the same clade with <i>Arabidopsis</i> genes but have not been investigated so far as flowering genes ... Arrows show promoting effects, T-bars show repressing effects. Environmental cues are shown as lower case letters in square boxes; ‘v’ is extended cold (vernalization); ‘ld’ is long days; ‘sd’ is short days; ‘am’ is ambient (non-vernalizing) temperature. Genes are shown in italics and proteins in non-italics in ovals. ‘Pfr’ indicates Pfr phytochrome signaling. <i>Arabidopsis</i> genes assigned to specific pathways are color-coded (photoperiod pathway in green, vernalization in blue and autonomous pathway in purple). Flowering pathway integrators are shown in red. Triple headed arrows indicate activation by red or blue light."
| |
| − | <br/>Overall pathway for the time of bloom from [http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0038250 Jung et al., 2012].
| |
| | | | |
| − | ===Additional <i>Arabidopsis</i> Flowering Pathways=== | + | ====Autonomous Pathway==== |
| | + | *The autonomous pathway is "activated in response to endogenous changes that are independent from the environmental cues leading to flowering", such as the plant's circadian rhythm ([http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0038250 Jung ''et al.'', 2012]) |
| | + | *17 <i>Arabidopsis</i> genes are involved in the autonomous pathway: CLF, FCA, FIE1, FLD, FLK, FPA, FVE, FY, LD, MSI1, SWN, VEL1, VEL2, VEL3, VIN3, VRN2, VRN5 ([http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0038250 Jung ''et al.'', 2012]) |
| | + | |
| | + | ====Gibberellin Pathway==== |
| | + | *Gibberellin "is essential for floral induction in short-day conditions." In fact, plants with a "mutation in a GA biosynthetic gene, such as GA1, fail to flower" ([http://www.plantcell.org/content/early/2012/08/29/tpc.112.101014.full.pdf+html Yu ''et al.'', 2012]) |
| | + | *5 <i>Arabidopsis</i> genes are involved in the gibberellin pathway: GAI, GID1, RGA, RGL1, RGL2 ([http://www.plantcell.org/content/early/2012/08/29/tpc.112.101014.full.pdf+html Yu ''et al.'', 2012]) |
| | + | |
| | + | ====Light Signaling Pathway==== |
| | + | *"Light is one of the main environmental regulators of flowering in plants. Plants sense the time of day and season of year by monitoring the light environment through light signalling pathways." Furthermore, the light signalling pathway is comprised of the "photoperiod pathway genes together with photoreceptor genes and circadian clock components" ([http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0038250 Jung ''et al.'', 2012]) |
| | + | *48 <i>Arabidopsis</i> genes are involved in the light signaling pathway: APRR3, APRR5, APRR9, AT1G26790, AT1G29160, AT2G34140, AT3G21320, AT3G25730, ATCOL4, ATCOL5, CCA1, CDF1, CDF2, CDF3, CDF5, CHE, CIB1, CO, COL1, COL2, COL9, COP1, CRY1, CRY2, ELF3, ELF4, ELF4-L3, FKF1, GI, LHY, LKP2, LUX, PHYA, PHYB, PHYC, PHYD, PHYE, PRR7, RAV1, RFI2, SPA1, SPA2, SPA3, SPA4, TEM1, TEM2, TOC1, ZTL ([http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0038250 Jung ''et al.'', 2012]) |
| | + | |
| | + | ====Polycomb Pathway==== |
| | + | *The polycomb pathway centers on “epigenetic [repression] … [of] various developmental and cellular processes … [through two] multi-subunit protein complexes: Polycomb Repressor Complex 1 (PRC1)” and Polycomb Repressor Complex 2 (PRC2) ([http://www.plosgenetics.org/article/info%3Adoi%2F10.1371%2Fjournal.pgen.1002512 Kim ''et al.'', 2012]) |
| | + | *10 <i>Arabidopsis</i> genes are involved in the polycomb pathway: CLF, EMF1, EMF2, FIE1, FIS2, LHP1, MEA, MSI1, SWN, VRN2 ([http://www.plosgenetics.org/article/info%3Adoi%2F10.1371%2Fjournal.pgen.1002512 Kim ''et al.'', 2012]) |
| | + | |
| | + | ====Vernalization Pathway==== |
| | + | *The vernalization pathway is the response to "prolonged periods of low temperature [that are required] to initiate flowering" ([http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0038250 Jung ''et al.'', 2012]) |
| | + | *32 <i>Arabidopsis</i> genes are involved in the vernalization pathway: AGL14, AGL19, AGL24, AGL31, ATARP6, ATSWC6, CLF, EFS, FES1, FIE1, FLC, FRI, FRL1, FRL2, HUA2, MAF1, MAF3, MAF4, MAF5, MSI1, PAF1, PAF2, PEP, PIE1, SUF4, SVP, SWN, VEL1, VIN3, VRN1, VRN2, VRN5 ([http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0038250 Jung ''et al.'', 2012]) |
| | + | |
| | + | |
| | + | ==Gallery of <i>Arabidopsis</i> Flowering Pathways== |
| | + | |
| | + | <gallery widths="102px" heights="102px" perrow="6"> |
| | + | File:Izawa_Pathway.jpeg|alt=Diagram of pathway from Izawa ''et al.'', 2003|[http://www.sciencedirect.com/science/article/pii/S1369526603000141 Izawa ''et al.'', 2003]; permission pending |
| | + | File:Sung_Pathway.gif|alt=Diagram of pathway from Sung ''et al.'', 2003|[http://www.sciencedirect.com/science/article/pii/S1369526602000146 Sung ''et al.'', 2003]; permission granted |
| | + | File:Boss_Pathway.gif|alt=Diagram of pathway from |[http://www.plantcell.org/content/16/suppl_1/S18/F2.expansion Boss ''et al.'', 2004]; permission granted |
| | + | File:Boss_Pathway_2.jpeg|alt=Diagram of pathway from Boss ''et al.'', 2004|[http://www.plantcell.org/content/16/suppl_1/S18/F3.expansion Boss ''et al.'', 2004]; permission granted |
| | + | File:Henderson_and_Dean_Pathway.jpeg|alt=Diagram of pathway from Henderson and Dean, 2004|[http://dev.biologists.org/content/131/16/3829/F1.expansion.html Henderson and Dean, 2004]; permission granted |
| | + | File:Amasino_Pathway.jpeg|alt=Diagram of pathway from Amasino, 2005|[http://www.sciencedirect.com/science/article/pii/S0958166905000273 Amasino, 2005]; permission granted |
| | + | File:He_and_Amasino_Pathway.jpeg|alt=Diagram of pathway from |[http://www.sciencedirect.com/science/article/pii/S1360138504002705 He and Amasino, 2005]; permission granted |
| | + | File:Yamaguchi_Pathway.jpeg|alt=Diagram of pathway from He and Amasino, 2005 |[http://pcp.oxfordjournals.org/content/46/8/1175/F7.expansion Yamaguchi ''et al.'', 2005]; permission pending |
| | + | File:Baurle_and_Dean_Pathway.jpeg|alt=Diagram of pathway from Yamaguchi ''et al.'', 2005|[http://www.sciencedirect.com/science/article/pii/S009286740600571X Bäurle and Dean, 2006]; permission granted |
| | + | File:Farrona_Pathway.jpeg|alt=Diagram of pathway from Farrona ''et al.'', 2008 |[http://www.sciencedirect.com/science/article/pii/S1084952108000566 Farrona ''et al.'', 2008]; permission granted |
| | + | File:Farrona_Pathway_2.jpeg|alt=Diagram of pathway from |[http://www.sciencedirect.com/science/article/pii/S1084952108000566 Farrona ''et al.'', 2008]; permission granted |
| | + | File:Amasino_Pathway_2.png|alt=Diagram of pathway from Amasino, 2010|[http://onlinelibrary.wiley.com/doi/10.1111/j.1365-313X.2010.04148.x/pdf/ Amasino, 2010]; permission granted |
| | + | File:Higgins_Pathway.jpg|alt=Diagram of pathway from Higgins ''et al.'', 2010 |[http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0010065 Higgins ''et al.'', 2010]; permission granted |
| | + | File:Kim_and_Sung_Pathway.jpeg|alt=Pathway from Kim and Sung, 2010 |[http://www.pnas.org/content/107/39/17029/F6.expansion.html Kim and Sung, 2010]; permission pending |
| | + | File:Taiz_and_Zeiger_Pathway.jpeg|alt=Diagram of pathway from Taiz and Zeiger, 2010 |[http://5e.plantphys.net/article.php?ch=1&id=375 Taiz and Zeiger, 2010]; permission pending |
| | + | File:Ballerini_and_Kramer_Pathway.png|alt=Diagram of pathway from Ballerini and Kramer, 2011 |[http://dash.harvard.edu/bitstream/handle/1/4903810/Ballerini%20%26%20Kramer%202011.pdf?sequence=1 Ballerini and Kramer, 2011]; permission granted |
| | + | File:Ferrier_Pathway.jpeg|alt=Diagram of pathway from |[http://www.sciencedirect.com/science/article/pii/S0958166910002284 Ferrier ''et al.'', 2011]; permission pending |
| | + | File:Zhang_Pathway.png|alt=Diagram of pathway from Ferrier ''et al.'', 2011|[http://www.biomedcentral.com/1471-2164/12/63 Zhang ''et al.'', 2011]; permission granted |
| | + | File:Jung_Pathway.jpg|alt=Diagram of pathway from Jung ''et al.'', 2012 |[http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0038250 Jung ''et al.'', 2012]; permission pending |
| | + | </gallery> |
| | + | |
| | + | Additional figures: |
| | + | *[http://dev.biologists.org/content/136/20/3379/F3.expansion.html Figure 3 from Liu ''et al.'', 2009] |
| | + | *[http://www.els.net/WileyCDA/ElsArticle/refId-a0002053.html Figure 1 from Schneitz and Balasubramanian, 2009] |
| | + | *[http://www.sciencedirect.com/science/article/pii/S0168952510001873 Figure 1 from Wellmer and Riechmann, 2010] |
| | + | *[http://www.sciencedirect.com/science/article/pii/S1369526611001440 Figure 1 from Posé ''et al.'', 2012] |
| | | | |
| − | {| class="table"
| |
| − | |-
| |
| − | | [[Image:Boss_Pathway.gif|110px|thumbnail|none|[http://www.plantcell.org/content/16/suppl_1/S18/F2.expansion Boss et al., 2004]]]||[[Image:Yamaguchi_Pathway.jpeg|110px|thumbnail|none|[http://pcp.oxfordjournals.org/content/46/8/1175/F7.expansion Yamaguchi et al., 2005]]]|| [[Image:Liu_Pathway.jpeg|110px|thumbnail|none|[http://dev.biologists.org/content/136/20/3379/F3.expansion.html Liu et al., 2009]]]||[[Image:Kim_and_Sung_Pathway.jpeg|110px|thumbnail|none|[http://www.pnas.org/content/107/39/17029/F6.expansion.html Kim and Sung, 2010]]]||[[Image:Taiz_and_Zeiger_Pathway.jpeg|110px|thumbnail|none|[http://5e.plantphys.net/article.php?ch=1&id=375 Taiz and Zeiger, 2010]]]||[[Image:Ballerini_and_Kramer_Pathway.png|110px|thumbnail|none|[http://openi.nlm.nih.gov/detailedresult.php?img=3049749_2041-9139-2-4-1&query=the&fields=all&favor=none&it=none&sub=none&uniq=0&sp=none&req=4&simCollection=1187896_1471-2202-6-48-3&npos=55&prt=3 Ballerini and Kramer, 2011]]]||[[Image:Zhang_Pathway.png|110px|thumbnail|none|[http://openi.nlm.nih.gov/detailedresult.php?img=3039610_1471-2164-12-63-6&query=the&fields=all&favor=none&it=none&sub=none&uniq=0&sp=none&req=4&simResults=f0a0c433%20f0a1c418%20f0a2c0%20f1a0c161%20f2a0c444%20f2a1c438%20f3a0c224%20f4a0c18%20f4a1c1%20f4a2c23&npos=37&prt=2 Zhang et al., 2011]]]||
| |
| − | |-
| |
| − | | [[Image:Pose_Pathway.jpeg|110px|thumbnail|none|[http://www.sciencedirect.com/science/article/pii/S1369526611001440 Posé et al., 2012]]]||
| |
| − | |}
| |
| | | | |
| | + | ==Methods== |
| | + | ====Finding Genes==== |
| | + | *I examined a variety of journal articles related to time of flowering in <i>Arabidopsis thaliana</i> and found a number of pathways related to flowering (see the gallery above). I came across a genomic analysis of soybean by [http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0038250 Jung ''et al.'', 2012]. In this paper, they listed the "183 <i>Arabidopsis</i> genes that are known to take part in flowering regulatory pathways [taken] from previous studies." These 183 genes, plus "24 additional <i>Arabidopsis</i> genes that are grouped into the same [homolog groups] as known flowering genes," provided a solid foundation for my study. All 207 total genes from [http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0038250 Jung ''et al.'', 2012] can be viewed here: [[File:Jung 207 Arabidopsis Flowering Genes.pdf]]. |
| | + | *These 207 total genes fell into two categories: 1) flowering pathway integrators/meristem identity genes and 2) condition pathway genes (responding to the photoperiod pathway, the vernalization pathway, the ambient temperature pathway, the autonomous pathway, and other pathways). Per the direction of Dr. Jeannie Rowland of the USDA Genetic Improvement for Fruits and Vegetables Laboratory, I focused on the condition pathway genes. |
| | + | *I identified a total of seven different pathways controlling flowering: the age pathway, the ambient temperature pathway, the autonomous pathway, the gibberellin pathway, the light signaling pathway, the polycomb pathway, and the vernalization pathway. Descriptions and primary genes involved in these pathways were taken from [http://onlinelibrary.wiley.com/doi/10.1111/j.1365-313X.2010.04148.x/pdf/ Amasino, 2010], [http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0038250 Jung ''et al.'', 2012], [http://www.plosgenetics.org/article/info%3Adoi%2F10.1371%2Fjournal.pgen.1002512 Kim ''et al.'', 2012], and [http://www.plantcell.org/content/early/2012/08/29/tpc.112.101014.full.pdf+html Yu ''et al.'', 2012]. |
| | + | *A total of 108 genes were examined, almost all of which "have been implicated in flowering-time control based on isolation of loss-of-function mutations or analysis of transgenic plants." ([http://www.sciencedirect.com/science/article/pii/S0092867410004411 Fornara ''et al.'', 2010]) |
| | | | |
| − | [http://www.sciencedirect.com/science/article/pii/S1084952108000566 Farrona et al., 2008]; | + | ====Finding SSRs==== |
| − | [http://www.sciencedirect.com/science/article/pii/S1360138504002705 He and Amasino, 2005];
| + | *A local database of the blueberry genome was created using the following coding: |
| − | [http://www.mpipz.mpg.de/305695/Project_1 Max Planck Institute for Plant Breeding Research];
| + | <PRE>./bin/makeblastdb -in BlueberryGenome.txt -input_type fasta -dbtype nucl -title blueberry_Genome</PRE> |
| − | [http://www.sciencedirect.com/science/article/pii/S009286740600571X Bäurle and Dean, 2006];
| + | *Amino acid sequences for all <i>Arabidopsis</i> genes were taken from [http://www.arabidopsis.org/index.jsp The Arabidopsis Information Resource (TAIR) (Huala ''et al.'', 2001)]. The amino acid sequences were run via tBLASTn against the blueberry scaffolds to find the closest match using the following coding: |
| − | [http://www.sciencedirect.com/science/article/pii/S1369526602000146 Sung et al., 2003];
| + | <PRE>{ echo bin/tblastn -query AASequence.txt -db BlueberryGenome.txt; bin/tblastn -query |
| − | [http://www.sciencedirect.com/science/article/pii/S0958166905000273 Amasino, 2005];
| + | AASequence.txt -db BlueberryGenome.txt; } >> AAOutput.txt</PRE> |
| − | [http://www.sciencedirect.com/science/article/pii/S1369526603000141 Izawa et al., 2003]
| + | *For each gene result, the best match was presumed to be the ortholog of the <i>Arabidopsis</i> gene in <i>Vaccinium corymbosum</i>. A maximum E value cutoff of e-04 was established. Although all of the results fell within this cutoff, if a tBLASTn result had not fallen below the E value limit, attempts would have been made to find and tBLASTn a <i>Vitis vinifera</i> ortholog of the <i>Arabidopsis</i> gene from [http://www.uniprot.org/uniprot/?query=organism%3A%22Vitis+vinifera+%5B29760%5D%22&sort=score UniProtKB (UniProt Consortium, 2012)] nomenclature search. |
| | + | *SSRs were determined by importing the best match scaffold into the [http://www.vaccinium.org/cgi-bin/vaccinium_ssr SSR Tool] at the [http://www.vaccinium.org/ Genome Database for Vaccinium]. Three di/trinucleotide SSRs near the gene location on the scaffold were chosen for each gene. |
| | | | |
| − | ===Core <i>Arabidopsis</i> Flowering Genes===
| |
| | | | |
| − | {| class="wikitable"
| + | ==SSR Results== |
| − | |-
| + | All SSR results can be viewed on the [[Time of bloom SSR Results]] page. An abbreviated listing of the results is included below. |
| − | ! <i>Arabidopsis</i> Gene !! Function (All function information is directly copied from the respective AA Sequence website link) !! AA Sequence !! Pathway !! Potential Ortholog
| + | <br/>[[File:Flowering_SSR_Results_Table.png]] |
| − | |-
| |
| − | | AG||Probable transcription factor involved in the control of organ identity during the early development of flowers. Is required for normal development of stamens and carpels in the wild-type flower. Plays a role in maintaining the determinacy of the floral meristem. Acts as C class cadastral protein by repressing the A class floral homeotic genes like APETALA1. Forms a heterodimer via the K-box domain with either SEPALATTA1/AGL2, SEPALATTA2/AGL4, SEPALLATA3/AGL9 or AGL6 that could be involved in genes regulation during floral meristem development.||[http://www.uniprot.org/uniprot/P17839 UniProtKB]||||[http://www.uniprot.org/uniprot/D1MDP5 Grape]
| |
| − | |-
| |
| − | | AGL24||Transcription activator that mediates floral transition in response to vernalization. Promotes inflorescence fate in apical meristems. Acts in a dosage-dependent manner. Probably involved in the transduction of RLK-mediated signaling (e.g. IMK3 pathway). Together with AP1 and SVP, controls the identity of the floral meristem and regulates expression of class B, C and E genes. When associated with SOC1, mediates effect of gibberellins on flowering under short-day conditions, and regulates the expression of LEAFY (LFY), which links floral induction and floral development. Confers inflorescence characteristics to floral primordia and early flowering.||[http://www.uniprot.org/uniprot/O82794 UniProtKB]||V||
| |
| − | |-
| |
| − | | AGL31||Probable transcription factor that prevents vernalization by short periods of cold. Acts as a floral repressor.||[http://www.uniprot.org/uniprot/Q9FPN7 UniProtKB]||Am, FPI, V||
| |
| − | |-
| |
| − | | AP1||Transcription factor that promotes early floral meristem identity in synergy with LEAFY. Is required subsequently for the transition of an inflorescence meristem into a floral meristem. Is indispensable for normal development of sepals and petals in flowers. Regulates positively the B class homeotic proteins APETALA3 and PISTILLATA with the cooperation of LEAFY and UFO. Interacts with SEPALLATA3 or AP3/PI heterodimer to form complexes that could be involved in genes regulation during floral meristem development. Regulates positively AGAMOUS in cooperation with LEAFY. Displays a redundant function with CAULIFLOWER in the up-regulation of LEAFY. Together with AGL24 and SVP, controls the identity of the floral meristem and regulates expression of class B, C and E genes. Represses flowering time genes AGL24, SVP and SOC1 in emerging floral meristems.||[http://www.uniprot.org/uniprot/P35631 UniProtKB]||FPI, MI||[http://www.uniprot.org/uniprot/D1MDP8 Grape]
| |
| − | |-
| |
| − | | AP2||Probable transcriptional activator that promotes early floral meristem identity. Is required subsequently for the transition of an inflorescence meristem into a floral meristem. Plays a central role in the specification of floral identity, particularly for the normal development of sepals and petals in the wild-type flower. Acts as A class cadastral protein by repressing the C class floral homeotic gene AGAMOUS in association with other repressors like LEUNIG and SEUSS. It is also required during seed development.||[http://www.uniprot.org/uniprot/P47927 UniProtKB]||||[http://www.uniprot.org/uniprot/C1KBJ2 Grape]
| |
| − | |-
| |
| − | | AP3||Probable transcription factor involved in the genetic control of flower development. Is required for normal development of petals and stamens in the wild-type flower. Forms a heterodimer with PISTILLATA that is required for autoregulation of both AP3 and PI genes. AP3/PI heterodimer interacts with APETALA1 or SEPALLATA3 to form a ternary complex that could be responsible for the regulation of the genes involved in the flower development. AP3/PI heterodimer activates the expression of NAP.||[http://www.uniprot.org/uniprot/P35632 UniProtKB]||||[http://www.uniprot.org/uniprot/A3RJI1 Grape]
| |
| − | |-
| |
| − | | CAL||Probable transcription factor that promotes early floral meristem identity in synergy with APETALA1, FRUITFULL and LEAFY. Is required subsequently for the transition of an inflorescence meristem into a floral meristem. Seems to be partially redundant to the function of APETALA1.||[http://www.uniprot.org/uniprot/Q39375 UniProtKB]||FPI, MI||
| |
| − | |-
| |
| − | | CO||Encodes a protein showing similarities to zinc finger transcription factors, involved in regulation of flowering under long days. Acts upstream of FT and SOC1.||[http://arabidopsis.org/servlets/TairObject?id=130492&type=locus TAIR]||FPI, L||
| |
| − | |-
| |
| − | | EMF1||Involved in regulating reproductive development||[http://arabidopsis.org/servlets/TairObject?id=130620&type=locus TAIR]||||
| |
| − | |-
| |
| − | | EMF2||Polycomb group protein with zinc finger domain involved in negative regulation of reproductive development. Forms a complex with FIE, CLF, and MSI1. This complex modulates the expression of target genes including AG, PI and AP3.||[http://arabidopsis.org/servlets/TairObject?id=134952&type=locus TAIR]||||
| |
| − | |-
| |
| − | | FCA||Involved in the promotion of the transition of the vegetative meristem to reproductive development. Four forms of the protein (alpha, beta, delta and gamma) are produced by alternative splicing. Involved in RNA-mediated chromatin silencing. At one point it was believed to act as an abscisic acid receptor but the paper describing that function was retracted.||[http://arabidopsis.org/servlets/TairObject?id=128853&type=locus TAIR]||Am, Au||[http://www.uniprot.org/uniprot/D2Y3W8 Grape]
| |
| − | |-
| |
| − | | FD||bZIP protein required for positive regulation of flowering. Mutants are late flowering. FD interacts with FT to promote flowering. Expressed in the shoot apex in floral anlagen, then declines in floral primordia.||[http://arabidopsis.org/servlets/TairObject?id=128068&type=locus TAIR]||Am, MI||
| |
| − | |-
| |
| − | | FLC||MADS-box protein encoded by FLOWERING LOCUS C - transcription factor that functions as a repressor of floral transition and contributes to temperature compensation of the circadian clock. Expression is downregulated during cold treatment. Vernalization, FRI and the autonomous pathway all influence the state of FLC chromatin. Both maternal and paternal alleles are reset by vernalization, but their earliest activation differs in timing and location. Histone H3 trimethylation at lysine 4 and histone acetylation are associated with active FLC expression, whereas histone deacetylation and histone H3 dimethylation at lysines 9 and 27 are involved in FLC repression. Expression is also repressed by two small RNAs (30- and 24-nt) complementary to the FLC sense strand 3’ to the polyA site. The small RNAs are most likely derived from an antisense transcript of FLC. Interacts with SOC1 and FT chromatin in vivo. Member of a protein complex.||[http://arabidopsis.org/servlets/TairObject?id=136002&type=locus TAIR]||Am, FPI, V||[http://www.uniprot.org/uniprot/D1MDP4 Grape]
| |
| − | |-
| |
| − | | FLD||Encodes a plant homolog of a SWIRM domain containing protein found in histone deacetylase complexes in mammals. Lesions in FLD result in hyperacetylation of histones in FLC chromatin, up-regulation of FLC expression and extremely delayed flowering. FLD plays a key role in regulating the reproductive competence of the shoot and results in different developmental phase transitions in Arabidopsis.||[http://arabidopsis.org/servlets/TairObject?id=35869&type=locus TAIR]||Am, Au||
| |
| − | |-
| |
| − | | FLK||RNA binding, nucleic acid binding; positive regulation of flower development||[http://arabidopsis.org/servlets/TairObject?id=37455&type=locus TAIR]||Au||
| |
| − | |-
| |
| − | | FPA||FPA is a gene that regulates flowering time in Arabidopsis via a pathway that is independent of daylength (the autonomous pathway). Mutations in FPA result in extremely delayed flowering. Double mutants with FCA have reduced fertility and single/double mutants have defects in siRNA mediated chromatin silencing.||[http://arabidopsis.org/servlets/TairObject?id=34279&type=locus TAIR]||Au||
| |
| − | |-
| |
| − | | FRI||Encodes a major determinant of natural variation in Arabidopsis flowering time. Dominant alleles of FRI confer a vernalization requirement causing plants to overwinter vegetatively. Many early flowering accessions carry loss-of-function fri alleles .Twenty distinct haplotypes that contain non-functional FRI alleles have been identified and the distribution analyzed in over 190 accessions. The common lab strains- Col and Ler each carry loss of function mutations in FRI.||[http://arabidopsis.org/servlets/TairObject?id=128299&type=locus TAIR]||V||
| |
| − | |-
| |
| − | | FRL1||Encodes a sterol-C24-methyltransferases involved in sterol biosynthesis. Mutants display altered sterol composition, serrated petals and sepals and altered cotyledon vascular patterning as well as ectopic endoreduplication. This suggests that suppression of endoreduplication is important for petal morphogenesis and that normal sterol composition is required for this suppression.||[http://arabidopsis.org/servlets/TairObject?id=27665&type=locus TAIR]||V||
| |
| − | |-
| |
| − | | FRL2||Family member of FRI-related genes that is required for the winter-annual habit. ||[http://arabidopsis.org/servlets/TairObject?id=226970&type=locus TAIR]||V||
| |
| − | |-
| |
| − | | FT||FT, together with LFY, promotes flowering and is antagonistic with its homologous gene, TERMINAL FLOWER1 (TFL1). FT is expressed in leaves and is induced by long day treatment. Either the FT mRNA or protein is translocated to the shoot apex where it induces its own expression. Recent data suggests that FT protein acts as a long-range signal. FT is a target of CO and acts upstream of SOC1.||[http://arabidopsis.org/servlets/TairObject?id=30541&type=locus TAIR]||Am, FPI||[http://www.uniprot.org/uniprot/D1MDP3 Grape], [http://www.uniprot.org/uniprot/E5BBS7 Strawberry]
| |
| − | |-
| |
| − | | FUL||MADS box gene negatively regulated by APETALA1||[http://arabidopsis.org/servlets/TairObject?id=134558&type=locus TAIR]||MI||[http://www.uniprot.org/uniprot/D1MDP9 Grape]
| |
| − | |-
| |
| − | | FVE||Controls flowering.||[http://arabidopsis.org/servlets/TairObject?id=33019&type=locus TAIR]||Am, Au||
| |
| − | |-
| |
| − | | FY||Encodes a protein with similarity to yeast Pfs2p, an mRNA processing factor. Involved in regulation of flowering time; affects FCA mRNA processing. Homozygous mutants are late flowering, null alleles are embryo lethal.||[http://arabidopsis.org/servlets/TairObject?id=136108&type=locus TAIR]||Au||
| |
| − | |-
| |
| − | | GI||Together with CONSTANTS (CO) and FLOWERING LOCUS T (FT), GIGANTEA promotes flowering under long days in a circadian clock-controlled flowering pathway. GI acts earlier than CO and FT in the pathway by increasing CO and FT mRNA abundance. Located in the nucleus. Regulates several developmental processes, including photoperiod-mediated flowering, phytochrome B signaling, circadian clock, carbohydrate metabolism, and cold stress response. The gene's transcription is controlled by the circadian clock and it is post-transcriptionally regulated by light and dark. Forms a complex with FKF1 on the CO promoter to regulate CO expression.||[http://arabidopsis.org/servlets/TairObject?id=136959&type=locus TAIR]||L||
| |
| − | |-
| |
| − | | LD||Encodes a nuclear localized protein with similarity to transcriptional regulators. Recessive mutants are late flowering. Expression of LFY is reduced in LD mutants.||[http://arabidopsis.org/servlets/TairObject?id=26564&type=locus TAIR]||Au||
| |
| − | |-
| |
| − | | LFY||Encodes transcriptional regulator that promotes the transition to flowering. Involved in floral meristem development. LFY is involved in the regulation of AP3 expression, and appears to bring the F-box protein UFO to the AP3 promoter. Amino acids 46-120 define a protein domain that mediates self-interaction.||[http://arabidopsis.org/servlets/TairObject?id=132616&type=locus TAIR]||FPI, MI||[http://www.uniprot.org/uniprot/Q8L5W6 Grape], [http://www.uniprot.org/uniprot/C4NF85 Strawberry]
| |
| − | |-
| |
| − | | MAF1||MADS domain protein - flowering regulator that is closely related to FLC. Deletion of this locus in Nd ecotype is correlated with earlier flowering in short days suggesting function as a negative regulator of flowering.||[http://arabidopsis.org/servlets/TairObject?id=29106&type=locus TAIR]||Am, FPI, V||
| |
| − | |-
| |
| − | | PI||Floral homeotic gene encoding a MADS domain transcription factor. Required for the specification of petal and stamen identities.||[http://arabidopsis.org/servlets/TairObject?id=131273&type=locus TAIR]||||[http://www.uniprot.org/uniprot/J9SQF2 Grape]
| |
| − | |-
| |
| − | | PIE1||Encodes a protein similar to ATP-dependent, chromatin-remodeling proteins of the ISWI and SWI2/SNF2 family. Genetic analyses suggest that this gene is involved in multiple flowering pathways. Mutations in PIE1 results in suppression of FLC-mediated delay of flowering and causes early flowering in noninductive photoperiods independently of FLC. PIE1 is required for expression of FLC in the shoot apex but not in the root. Along with ARP6 forms a complex to deposit modified histone H2A.Z at several loci within the genome. This modification alters the expression of the target genes (i.e. FLC, MAF4, MAF6).||[http://arabidopsis.org/servlets/TairObject?id=37964&type=locus TAIR]||V||
| |
| − | |-
| |
| − | | SMZ||Encodes a AP2 domain transcription factor that can repress flowering. SMZ and its paralogous gene, SNARCHZAPFEN (SNZ), share a signature with partial complementarity to the miR172 microRNA, whose precursor is induced upon flowering.||[http://arabidopsis.org/servlets/TairObject?id=37065&type=locus TAIR]||||
| |
| − | |-
| |
| − | | SNZ||Encodes a AP2 domain transcription factor that can repress flowering. SNZ and its paralogous gene, SCHLAFMUTZE (SMZ), share a signature with partial complementarity to the miR172 microRNA, whose precursor is induced upon flowering.||[http://arabidopsis.org/servlets/TairObject?id=33943&type=locus TAIR]||||
| |
| − | |-
| |
| − | | SOC1||Transcription activator active in flowering time control. May integrate signals from the photoperiod, vernalization and autonomous floral induction pathways. Can modulate class B and C homeotic genes expression. When associated with AGL24, mediates effect of gibberellins on flowering under short-day conditions, and regulates the expression of LEAFY (LFY), which links floral induction and floral development.||[http://www.uniprot.org/uniprot/O64645 UniProtKB]||FPI||[http://www.uniprot.org/uniprot/D1MDP7 Grape], [http://www.uniprot.org/uniprot/G3GAX1 Strawberry]
| |
| − | |-
| |
| − | | SPL3||Encodes a member of the SPL (squamosa-promoter binding protein-like)gene family, a novel gene family encoding DNA binding proteins and putative transcription factors. Contains the SBP-box, which encodes the SBP-domain, required and sufficient for interaction with DNA. It binds DNA, may directly regulate AP1, and is involved in regulation of flowering and vegetative phase change. Its temporal expression is regulated by the microRNA miR156. The target site for the microRNA is in the 3'UTR.||[http://arabidopsis.org/servlets/TairObject?id=34194&type=locus TAIR]||MI||
| |
| − | |-
| |
| − | | SPY||Encodes a N-acetyl glucosamine transferase that may glycosylate other molecules involved in GA signaling. Contains a tetratricopeptide repeat region, and a novel carboxy-terminal region. SPY acts as both a repressor of GA responses and as a positive regulation of cytokinin signaling. SPY may be involved in reducing ROS accumulation in response to stress.||[http://arabidopsis.org/servlets/TairObject?id=36698&type=locus TAIR]||||
| |
| − | |-
| |
| − | | SVP||Encodes a nuclear protein that acts as a floral repressor and that functions within the thermosensory pathway. SVP represses FT expression via direct binding to the vCArG III motif in the FT promoter.||[http://arabidopsis.org/servlets/TairObject?id=31694&type=locus TAIR]||Am, V||[http://www.uniprot.org/uniprot/H9CTT8 Grape]
| |
| − | |-
| |
| − | | TEM1||Encodes a member of the RAV transcription factor family that contains AP2 and B3 binding domains. Involved in the regulation of flowering under long days. Loss of function results in early flowering. Overexpression causes late flowering and repression of expression of FT. Novel transcriptional regulator involved in ethylene signaling. Promoter bound by EIN3. EDF1 in turn, binds to promoter elements in ethylene responsive genes.||[http://arabidopsis.org/servlets/TairObject?id=30061&type=locus TAIR]||L||
| |
| − | |-
| |
| − | | TEM2||Rav2 is part of a complex that has been named `regulator of the (H+)-ATPase of the vacuolar and endosomal membranes' (RAVE)||[http://arabidopsis.org/servlets/TairObject?id=27570&type=locus TAIR]||L||
| |
| − | |-
| |
| − | | TFL1||Controls inflorescence meristem identity. Involved in the floral initiation process. Ortholog of the Antirrhinum gene CENTRORADIALIS (CEN). Involved in protein trafficking to the protein storage vacuole.||[http://arabidopsis.org/servlets/TairObject?id=131459&type=locus TAIR]||Am, FPI||[http://www.uniprot.org/uniprot/Q84MI7 Grape], [http://www.uniprot.org/uniprot/G5CJU3 Strawberry]
| |
| − | |-
| |
| − | | TOE1||Pathogenesis-related transcriptional factor/ERF, DNA-binding||[http://arabidopsis.org/servlets/TairObject?id=34025&type=locus TAIR]||||
| |
| − | |-
| |
| − | | TOE2||Sequence-specific DNA binding transcription factor activity; regulation of transcription, DNA-dependent||[http://arabidopsis.org/servlets/TairObject?id=133319&type=locus TAIR]||||
| |
| − | |-
| |
| − | | TOE3||Pathogenesis-related transcriptional factor/ERF, DNA-binding||[http://arabidopsis.org/servlets/TairObject?id=132134&type=locus TAIR]||||
| |
| − | |-
| |
| − | | TSF||Encodes a floral inducer that is a homolog of FT. Plants overexpressing this gene flower earlier than Col. Loss-of-function mutations flower later in short days. TSF and FT play overlapping roles in the promotion of flowering, with FT playing the dominant role. TSF sequences show extensive variation in different accessions and may contribute to quantitative variation in flowering time in these accessions. TSF has a complex pattern of spatial expression; it is expressed mainly in phloem and expression is regulated by daylength and vernalization.||[http://arabidopsis.org/servlets/TairObject?id=26553&type=locus TAIR]||Am, FPI||
| |
| − | |-
| |
| − | | VIN3||Encodes a plant homeodomain protein VERNALIZATION INSENSITIVE 3 (VIN3). In planta VIN3 and VRN2, VERNALIZATION 2, are part of a large protein complex that can include the polycomb group (PcG) proteins FERTILIZATION INDEPENDENT ENDOSPERM (FIE), CURLY LEAF (CLF), and SWINGER (SWN or EZA1). The complex has a role in establishing FLC (FLOWERING LOCUS C) repression during vernalization.||[http://arabidopsis.org/servlets/TairObject?id=134483&type=locus TAIR]||Au, V||
| |
| − | |-
| |
| − | | VIP3||The protein is composed of repeats of WD motif which is involved in protein complex formation. The gene is involved in flower timing and flower development. This gene is predicted to encode a protein with a DWD motif. It can bind to DDB1a in Y2H assays, and DDB1b in co-IP assays, and may be involved in the formation of a CUL4-based E3 ubiquitin ligase. Loss of gene function leads to a redistribution of H3K4me3 and K3K36me2 modifications within genes but not a change in the overall abundance of these modifications within chromatin.||[http://arabidopsis.org/servlets/TairObject?id=127868&type=locus TAIR]||||
| |
| − | |-
| |
| − | | VIP4||Encodes highly hydrophilic protein involved in positively regulating FLC expression. Mutants are early flowering and show a loss of FLC expression in the absence of cold.||[http://arabidopsis.org/servlets/TairObject?id=132654&type=locus TAIR]||||
| |
| − | |-
| |
| − | | VRN1||Required for vernalization. Essential for the complete repression of FLC in vernalized plants. Required for the methylation of histone H3||[http://arabidopsis.org/servlets/TairObject?id=37620&type=locus TAIR]||V||
| |
| − | |-
| |
| − | | VRN2||The VERNALIZATION2 (VRN2) gene mediates vernalization and encodes a nuclear-localized zinc finger protein with similarity to Polycomb group (PcG) proteins of plants and animals. In wild-type Arabidopsis, vernalization results in the stable reduction of the levels of the floral repressor FLC. In vrn2 mutants, FLC expression is downregulated normally in response to vernalization, but instead of remaining low, FLC mRNA levels increase when plants are returned to normal temperatures. VRN2 maintains FLC repression after a cold treatment, serving as a mechanism for the cellular memory of vernalization. Required for complete repression of FLC. Required for the methylation of histone H3||[http://arabidopsis.org/servlets/TairObject?id=228106&type=locus TAIR]||Au, V||
| |
| − | |}
| |
| | | | |
| − | ===205 <i>Arabidopsis</i> Flowering Genes===
| |
| | | | |
| − | Note: All <i>Arabidopsis</i> genes are compiled from [http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0038250 Jung et al., 2012]. All potential orthologs are found via [http://www.uniprot.org/uniprot/?query=taxonomy%3A%22Vitis+vinifera+%28Grape%29+%5B29760%5D%22&sort=score UniProt Grape] or [http://www.uniprot.org/uniprot/?query=taxonomy%3A%22Fragaria+vesca+%5B57918%5D%22&sort=score UniProt Strawberry] nomenclature search. Formatting is the same as [http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0038250 Jung et al., 2012] where a * indicates 1 of "24 <i>Arabidopsis</i> genes, which had not been previously investigated for their roles in flowering, [that] belong to [paralog groups] with known <i>Arabidopsis</i> flowering genes." Moreover, for pathways, "Am: Ambient temperature pathway, Au: Autonomous pathway, FPI: Flowering Pathway Intergrators, L: Light signaling pathway, MI: Meristem Identity, V: Vernalization pathway."
| + | ==Flowering Genes Of Interest== |
| | + | This table lists 108 genes involved in the age, ambient temperature, autonomous, gibberellin, light signaling, polycomb, and vernalization pathways, almost all of which "have been implicated in flowering-time control based on isolation of loss-of-function mutations or analysis of transgenic plants." ([http://www.sciencedirect.com/science/article/pii/S0092867410004411 Fornara ''et al.'', 2010]) |
| | | | |
| − | {| class="wikitable" | + | {| class="wikitable sortable" |
| − | |-
| |
| − | ! <i>Arabidopsis</i> Gene !! Function (All function information is directly copied from the respective AA Sequence website link) !! AA Sequence !! Pathway !! Potential Ortholog
| |
| − | |-
| |
| − | | ABF1||Binds to the ABA-responsive element (ABRE). Could participate in abscisic acid-regulated gene expression.||[http://www.uniprot.org/uniprot/Q9M7Q5 UniProtKB]||||
| |
| − | |-
| |
| − | | ABF2||Involved in ABA and stress responses and acts as a positive component of glucose signal transduction. Functions as transcriptional activator in the ABA-inducible expression of rd29B. Binds specifically to the ABA-responsive element (ABRE) of the rd29B gene promoter.||[http://www.uniprot.org/uniprot/Q9M7Q4 UniProtKB]||||
| |
| − | |-
| |
| − | | ABF3||Binds to the ABA-responsive element (ABRE). Mediates stress-responsive ABA signaling.||[http://www.uniprot.org/uniprot/Q9M7Q3 UniProtKB]||||
| |
| − | |-
| |
| − | | ABF4||Functions as transcriptional activator in the ABA-inducible expression of rd29B. Binds specifically to the ABA-responsive element (ABRE) of the rd29B gene promoter.||[http://www.uniprot.org/uniprot/Q9M7Q2 UniProtKB]||||
| |
| − | |-
| |
| − | | ABI5||Participates in ABA-regulated gene expression during seed development and subsequent vegetative stage by acting as the major mediator of ABA repression of growth. Binds to the embryo specification element and the ABA-responsive element (ABRE) of the Dc3 gene promoter and to the ABRE of the Em1 and Em6 genes promoters. Can also trans-activate its own promoter, suggesting that it is autoregulated. Plays a role in sugar-mediated senescence.||[http://www.uniprot.org/uniprot/Q9SJN0 UniProtKB]||||
| |
| − | |-
| |
| − | | AG||Probable transcription factor involved in the control of organ identity during the early development of flowers. Is required for normal development of stamens and carpels in the wild-type flower. Plays a role in maintaining the determinacy of the floral meristem. Acts as C class cadastral protein by repressing the A class floral homeotic genes like APETALA1. Forms a heterodimer via the K-box domain with either SEPALATTA1/AGL2, SEPALATTA2/AGL4, SEPALLATA3/AGL9 or AGL6 that could be involved in genes regulation during floral meristem development.||[http://www.uniprot.org/uniprot/P17839 UniProtKB]||||[http://www.uniprot.org/uniprot/D1MDP5 Grape]
| |
| − | |-
| |
| − | | AGL14||Probable transcription factor.||[http://www.uniprot.org/uniprot/Q38838 UniProtKB]||V||
| |
| − | |-
| |
| − | | AGL15||Transcription factor involved in the negative regulation of flowering, probably through the photoperiodic pathway. Acts as both an activator and a repressor of transcription. Binds DNA in a sequence-specific manner in large CArG motif 5'-CC (A/T)8 GG-3'. Participates probably in the regulation of programs active during the early stages of embryo development. Prevents premature perianth senescence and abscission, fruits development and seed desiccation. Stimulates the expression of at least DTA4, LEC2, FUS3, ABI3, AT4G38680/CSP2 and GRP2B/CSP4. Can enhance somatic embryo development in vitro.||[http://www.uniprot.org/uniprot/Q38847 UniProtKB]||||
| |
| − | |-
| |
| − | | AGL17||Probable transcription factor.||[http://www.uniprot.org/uniprot/Q38840 UniProtKB]||||
| |
| − | |-
| |
| − | | AGL18||Probable transcription factor involved in the negative regulation of flowering, probably through the photoperiodic pathway. Prevents premature flowering. Downstream regulator of a subset of the MIKC* MADS-controlled genes required during pollen maturation.||[http://www.uniprot.org/uniprot/Q9M2K8 UniProtKB]||||
| |
| − | |-
| |
| − | | AGL19||Probable transcription factor that promotes flowering, especially in response to vernalization by short periods of cold, in an FLC-inpedendent manner.||[http://www.uniprot.org/uniprot/O82743 UniProtKB]||V||
| |
| − | |-
| |
| − | | AGL21*||Probable transcription factor.||[http://www.uniprot.org/uniprot/Q9SZJ6 UniProtKB]||||
| |
| − | |-
| |
| − | | AGL24||Transcription activator that mediates floral transition in response to vernalization. Promotes inflorescence fate in apical meristems. Acts in a dosage-dependent manner. Probably involved in the transduction of RLK-mediated signaling (e.g. IMK3 pathway). Together with AP1 and SVP, controls the identity of the floral meristem and regulates expression of class B, C and E genes. When associated with SOC1, mediates effect of gibberellins on flowering under short-day conditions, and regulates the expression of LEAFY (LFY), which links floral induction and floral development. Confers inflorescence characteristics to floral primordia and early flowering.||[http://www.uniprot.org/uniprot/O82794 UniProtKB]||V||
| |
| − | |-
| |
| − | | AGL31||Probable transcription factor that prevents vernalization by short periods of cold. Acts as a floral repressor.||[http://www.uniprot.org/uniprot/Q9FPN7 UniProtKB]||Am, FPI, V||
| |
| | |- | | |- |
| − | | AGL6||Probable transcription factor. Forms a heterodimer via the K-box domain with AG, that could be involved in genes regulation during floral meristem development.||[http://www.uniprot.org/uniprot/P29386 UniProtKB]||||
| + | ! <i>Arabidopsis</i> Locus !! Other Names !! AA Source !! Pathway !! width="150px" |Top Hit ''Vaccinium'' Scaffold !! E Value |
| | |- | | |- |
| − | | AGL79||||[http://www.uniprot.org/uniprot/Q7X9H6 UniProtKB]|||| | + | | AT1G01060||LATE ELONGATED HYPOCOTYL, LATE ELONGATED HYPOCOTYL 1, LHY, LHY1||[http://arabidopsis.org/servlets/TairObject?id=137165&type=locus TAIR]||Light Signaling||Scaffold00140 (length 354209) at 234299||2e-19 |
| | |- | | |- |
| − | | AP1||Transcription factor that promotes early floral meristem identity in synergy with LEAFY. Is required subsequently for the transition of an inflorescence meristem into a floral meristem. Is indispensable for normal development of sepals and petals in flowers. Regulates positively the B class homeotic proteins APETALA3 and PISTILLATA with the cooperation of LEAFY and UFO. Interacts with SEPALLATA3 or AP3/PI heterodimer to form complexes that could be involved in genes regulation during floral meristem development. Regulates positively AGAMOUS in cooperation with LEAFY. Displays a redundant function with CAULIFLOWER in the up-regulation of LEAFY. Together with AGL24 and SVP, controls the identity of the floral meristem and regulates expression of class B, C and E genes. Represses flowering time genes AGL24, SVP and SOC1 in emerging floral meristems.||[http://www.uniprot.org/uniprot/P35631 UniProtKB]||FPI, MI||[http://www.uniprot.org/uniprot/D1MDP8 Grape] | + | | AT1G02580||EMB173, EMBRYO DEFECTIVE 173, FERTILIZATION INDEPENDENT SEED 1, FIS1, MEA, MEDEA, SDG5, SET DOMAIN-CONTAINING PROTEIN 5||[http://arabidopsis.org/servlets/TairObject?id=136450&type=locus TAIR]||Polycomb||Scaffold00354 (length 215005) at 64805||4e-17 |
| | |- | | |- |
| − | | AP2||Probable transcriptional activator that promotes early floral meristem identity. Is required subsequently for the transition of an inflorescence meristem into a floral meristem. Plays a central role in the specification of floral identity, particularly for the normal development of sepals and petals in the wild-type flower. Acts as A class cadastral protein by repressing the C class floral homeotic gene AGAMOUS in association with other repressors like LEUNIG and SEUSS. It is also required during seed development.||[http://www.uniprot.org/uniprot/P47927 UniProtKB]||||[http://www.uniprot.org/uniprot/C1KBJ2 Grape] | + | | AT1G04400||AT-PHH1, ATCRY2, CRY2, CRYPTOCHROME 2, FHA, PHH1||[http://arabidopsis.org/servlets/TairObject?id=28374&type=locus TAIR]||Light Signaling||Scaffold00649 (length 159319) at 28296||1e-119 |
| | |- | | |- |
| − | | AP3||Probable transcription factor involved in the genetic control of flower development. Is required for normal development of petals and stamens in the wild-type flower. Forms a heterodimer with PISTILLATA that is required for autoregulation of both AP3 and PI genes. AP3/PI heterodimer interacts with APETALA1 or SEPALLATA3 to form a ternary complex that could be responsible for the regulation of the genes involved in the flower development. AP3/PI heterodimer activates the expression of NAP.||[http://www.uniprot.org/uniprot/P35632 UniProtKB]||||[http://www.uniprot.org/uniprot/A3RJI1 Grape] | + | | AT1G09570||ELONGATED HYPOCOTYL 8, FAR RED ELONGATED 1, FAR RED ELONGATED HYPOCOTYL 2, FHY2, FRE1, HY8, PHYA, PHYTOCHROME A||[http://arabidopsis.org/servlets/TairObject?id=27545&type=locus TAIR]||Light Signaling||Scaffold03861 (length 7403) at 3771||0.0 |
| | |- | | |- |
| − | | APRR3||Controls photoperiodic flowering response. Component of the circadian clock. Controls the degradation of APRR1/TOC1 by the SCF(ZTL) complex. Expression of several members of the ARR-like family is controlled by circadian rhythm. The particular coordinated sequential expression of APRR9, APRR7, APRR5, APRR3 and APPR1 result to circadian waves that may be at the basis of the endogenous circadian clock.||[http://www.uniprot.org/uniprot/Q9LVG4 UniProtKB]||L|| | + | | AT1G13260||EDF4, ETHYLENE RESPONSE DNA BINDING FACTOR 4, RAV1, RELATED TO ABI3/VP1 1||[http://arabidopsis.org/servlets/TairObject?id=137721&type=locus TAIR]||Light Signaling||Scaffold00930 (length 110378) at 63069||1e-103 |
| | |- | | |- |
| − | | APRR5||Controls photoperiodic flowering response. Seems to be one of the component of the circadian clock. Expression of several members of the ARR-like family is controlled by circadian rhythm. The particular coordinated sequential expression of APRR9, APRR7, APRR5, APRR3 and APPR1 result to circadian waves that may be at the basis of the endogenous circadian clock.||[http://www.uniprot.org/uniprot/Q6LA42 UniProtKB]||L|| | + | | AT1G14920||GAI, GIBBERELLIC ACID INSENSITIVE, RESTORATION ON GROWTH ON AMMONIA 2, RGA2||[http://arabidopsis.org/servlets/TairObject?id=26612&type=locus TAIR]||Gibberellin||Scaffold01360 (length 81306) at 51382||0.0 |
| | |- | | |- |
| − | | APRR9||Controls photoperiodic flowering response. Seems to be one of the component of the circadian clock. Expression of several members of the ARR-like family is controlled by circadian rhythm. The particular coordinated sequential expression of APRR9, APRR7, APRR5, APRR3 and APPR1 result to circadian waves that may be at the basis of the endogenous circadian clock.||[http://www.uniprot.org/uniprot/Q8L500 UniProtKB]||L|| | + | | AT1G20330||COTYLEDON VASCULAR PATTERN 1, CVP1, FRILL1, FRL1, SMT2, STEROL METHYLTRANSFERASE 2||[http://arabidopsis.org/servlets/TairObject?id=27665&type=locus TAIR]||Vernalization||Scaffold02142 (length 46662) at 24743||9e-06 |
| | |- | | |- |
| − | | AREB3||Binds to the embryo specification element and the ABA-responsive element (ABRE) of the Dc3 gene promoter. Could participate in abscisic acid-regulated gene expression during seed development.||[http://www.uniprot.org/uniprot/Q9LES3 UniProtKB]|||| | + | | AT1G22770||FB, GI, GIGANTEA||[http://arabidopsis.org/servlets/TairObject?id=136959&type=locus TAIR]||Light Signaling||Scaffold00100 (length 346620) at 198329||0.0 |
| | |- | | |- |
| − | | AT1G26790||Transcription factor that binds specifically to a 5'-AA[AG]G-3' consensus core sequence.||[http://www.uniprot.org/uniprot/Q9LQX4 UniProtKB]||L|| | + | | AT1G25560||EDF1, ETHYLENE RESPONSE DNA BINDING FACTOR 1, TEM1, TEMPRANILLO 1||[http://arabidopsis.org/servlets/TairObject?id=30061&type=locus TAIR]||Light Signaling||Scaffold00930 (length 110378) at 63096||3e-99 |
| | |- | | |- |
| − | | AT1G29160||Transcription factor that binds specifically to a 5'-AA[AG]G-3' consensus core sequence. Acts as a negative regulator in the phytochrome-mediated light responses. Controls phyB-mediated end-of-day response and the phyA-mediated anthocyanin accumulation.||[http://www.uniprot.org/uniprot/P68350 UniProtKB]||L|| | + | | AT1G26790||||[http://arabidopsis.org/servlets/TairObject?id=137136&type=locus TAIR]||Light Signaling||Scaffold00079 (length 471015) at 69048||6e-33 |
| | |- | | |- |
| − | | AT1G50680||Probably acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. May be involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways.||[http://www.uniprot.org/uniprot/Q9C6P5 UniProtKB]|||| | + | | AT1G27370||SPL10, SQUAMOSA PROMOTER BINDING PROTEIN-LIKE 10||[http://arabidopsis.org/servlets/TairObject?id=28131&type=locus TAIR]||Age||Scaffold00127 (length 336847) at 165806||5e-33 |
| | |- | | |- |
| − | | AT1G51120||Probably acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. May be involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways.||[http://www.uniprot.org/uniprot/Q9C688 UniProtKB]|||| | + | | AT1G29160||||[http://arabidopsis.org/servlets/TairObject?id=29883&type=locus TAIR]||Light Signaling||Scaffold00270 (length 246371) at 215229||4e-50 |
| | |- | | |- |
| − | | AT2G23070||Casein kinases are operationally defined by their preferential utilization of acidic proteins such as caseins as substrates. The alpha chain contains the catalytic site. May act as an ectokinase that phosphorylates several extracellular proteins.||[http://www.uniprot.org/uniprot/O64816 UniProtKB]|||| | + | | AT1G30970||SUF4, SUPPRESSOR OF FRIGIDA4||[http://arabidopsis.org/servlets/TairObject?id=28096&type=locus TAIR]||Vernalization||Scaffold00348 (length 210978) at 75279||1e-15 |
| | |- | | |- |
| − | | AT2G23080||Casein kinases are operationally defined by their preferential utilization of acidic proteins such as caseins as substrates. The alpha chain contains the catalytic site. May act as an ectokinase that phosphorylates several extracellular proteins.||[http://www.uniprot.org/uniprot/O64817 UniProtKB]|||| | + | | AT1G31814||FRIGIDA LIKE 2, FRL2||[http://arabidopsis.org/servlets/TairObject?id=226970&type=locus TAIR]||Vernalization||Scaffold00289 (length 259591) at 150896||2e-19 |
| | |- | | |- |
| − | | AT2G25920||Uncharacterized protein||[http://www.uniprot.org/uniprot/O82305 UniProtKB]|||| | + | | AT1G47250||20S PROTEASOME ALPHA SUBUNIT F2, PAF2||[http://arabidopsis.org/servlets/TairObject?id=30983&type=locus TAIR]||Vernalization||Scaffold00528 (length 197283) at 110933||3e-64 |
| | |- | | |- |
| − | | AT2G34140||Transcription factor that binds specifically to a 5'-AA[AG]G-3' consensus core sequence.||[http://www.uniprot.org/uniprot/O22967 UniProtKB]||L|| | + | | AT1G53090||SPA1-RELATED 4, SPA4||[http://arabidopsis.org/servlets/TairObject?id=31031&type=locus TAIR]||Light Signaling||Scaffold00734 (length 158513) at 143450||3e-129 |
| | |- | | |- |
| − | | AT2G46670*||Putative uncharacterized protein||[http://www.uniprot.org/uniprot/Q683A6 UniProtKB]||L|| | + | | AT1G53160||FLORAL TRANSITION AT THE MERISTEM6, FTM6, SPL4, SQUAMOSA PROMOTER BINDING PROTEIN-LIKE 4||[http://arabidopsis.org/servlets/TairObject?id=27097&type=locus TAIR]||Age||Scaffold00062 (length 451336) at 412489||1e-15 |
| | |- | | |- |
| − | | AT3G21320||Uncharacterized protein||[http://www.uniprot.org/uniprot/Q5Q0C8 UniProtKB]||L|| | + | | AT1G65480||FLOWERING LOCUS T, FT||[http://arabidopsis.org/servlets/TairObject?id=30541&type=locus TAIR]||Ambient Temperature||Scaffold00357 (length 260075) at 58774||6e-35 |
| | |- | | |- |
| − | | AT3G25730||Probably acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. May be involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways.||[http://www.uniprot.org/uniprot/Q9LS06 UniProtKB]||L|| | + | | AT1G66350||RGA-LIKE 1, RGL, RGL1||[http://arabidopsis.org/servlets/TairObject?id=137244&type=locus TAIR]||Gibberellin||Scaffold00134 (length 346346) at 174707||0.0 |
| | |- | | |- |
| − | | AT4G16810||Putative uncharacterized protein||[http://www.uniprot.org/uniprot/O23521 UniProtKB]|||| | + | | AT1G68050||"FLAVIN-BINDING, KELCH REPEAT, F BOX 1", ADO3, FKF1||[http://arabidopsis.org/servlets/TairObject?id=137051&type=locus TAIR]||Light Signaling||Scaffold00110 (length 358202) at 120279||0.0 |
| | |- | | |- |
| − | | AT5G08230*||Probable transcrition factor that seems to be involved in mRNA processing.||[http://www.uniprot.org/uniprot/Q9LEY4 UniProtKB]||V|| | + | | AT1G68840||ATRAV2, EDF2, ETHYLENE RESPONSE DNA BINDING FACTOR 2, RAP2.8, RAV2, RELATED TO ABI3/VP1 2, RELATED TO AP2 8, TEM2, TEMPRANILLO 2||[http://arabidopsis.org/servlets/TairObject?id=27570&type=locus TAIR]||Light Signaling||Scaffold00930 (length 110378) at 63096||5e-102 |
| | |- | | |- |
| − | | AT5G10625*||Modulates the competence to flowering of apical meristems.||[http://www.uniprot.org/uniprot/Q9LXB5 UniProtKB]|||| | + | | AT1G77080||AGAMOUS-LIKE 27, AGL27, FLM, FLOWERING LOCUS M, MADS AFFECTING FLOWERING 1, MAF1||[http://arabidopsis.org/servlets/TairObject?id=29106&type=locus TAIR]||Ambient Temperature, Vernalization||Scaffold10765 (length 2447) at 744||3e-13 |
| | |- | | |- |
| − | | AT5G23280*||Transcription factor||[http://www.uniprot.org/uniprot/Q9FMX2 UniProtKB]||L|| | + | | AT1G77300||ASH1 HOMOLOG 2, ASHH2, CAROTENOID CHLOROPLAST REGULATORY1, CCR1, EARLY FLOWERING IN SHORT DAYS, EFS, LAZ2, LAZARUS 2, SDG8, SET DOMAIN GROUP 8||[http://arabidopsis.org/servlets/TairObject?id=136429&type=locus TAIR]||Vernalization||Scaffold00894 (length 114877) at 89213||1e-27 |
| | |- | | |- |
| − | | AT5G27220||Frigida-like protein||[http://www.uniprot.org/uniprot/O04650 UniProtKB]|||| | + | | AT2G01570||REPRESSOR OF GA, REPRESSOR OF GA1-3 1, RGA, RGA1||[http://arabidopsis.org/servlets/TairObject?id=26549&type=locus TAIR]||Gibberellin||Scaffold01360 (length 81306) at 51382||0.0 |
| | |- | | |- |
| − | | AT5G42910||Could participate in abscisic acid-regulated gene expression.||[http://www.uniprot.org/uniprot/Q9FMM7 UniProtKB]|||| | + | | AT2G06255||ELF4-L3, ELF4-LIKE 3||[http://arabidopsis.org/servlets/TairObject?id=500231553&type=locus TAIR]||Light Signaling||Scaffold00336 (length 230445) at 188137||1e-39 |
| | |- | | |- |
| − | | AT5G44080||Basic leucine zipper transcription factor||[http://www.uniprot.org/uniprot/Q9FNB9 UniProtKB]|||| | + | | AT2G17770||ATBZIP27, BASIC REGION/LEUCINE ZIPPER MOTIF 27, BZIP27, FD PARALOG, FDP||[http://arabidopsis.org/servlets/TairObject?id=227228&type=locus TAIR]||Ambient Temperature||Scaffold00367 (length 240396) at 113139||7e-17 |
| | |- | | |- |
| − | | AT5G48250*||Zinc finger protein Constans-like 10||[http://www.uniprot.org/uniprot/Q9LUA9 UniProtKB]||L|| | + | | AT2G18790||HY3, OOP1, OUT OF PHASE 1, PHYB, PHYTOCHROME B||[http://arabidopsis.org/servlets/TairObject?id=26548&type=locus TAIR]||Light Signaling||Scaffold00751 (length 152548) at 83698||0.0 |
| | |- | | |- |
| − | | AT5G59570*||Putative transcription factor||[http://www.uniprot.org/uniprot/Q9LTH4 UniProtKB]||L|| | + | | AT2G18870||VEL3, VERNALIZATION5/VIN3-LIKE 3, VIL4, VIN3-LIKE 4||[http://arabidopsis.org/servlets/TairObject?id=32249&type=locus TAIR]||Autonomous||Scaffold00396 (length 187979) at 73810||7e-19 |
| | |- | | |- |
| − | | AT5G62040||May form complexes with phosphorylated ligands by interfering with kinases and their effectors.||[http://www.uniprot.org/uniprot/Q9FIT4 UniProtKB]|||| | + | | AT2G18880||VEL2, VERNALIZATION5/VIN3-LIKE 2, VIL3, VIN3-LIKE 3||[http://arabidopsis.org/servlets/TairObject?id=32230&type=locus TAIR]||Autonomous||Scaffold00026 (length 499904) at 369396||6e-41 |
| | |- | | |- |
| − | | ATARP6||Component of the SWR1 complex which mediates the ATP-dependent exchange of histone H2A for the H2A variant H2A.F/Z leading to transcriptional regulation of selected genes (e.g. FLC) by chromatin remodeling. Required for the activation of FLC and FLC/MAF genes expression to levels that inhibit flowering, through both histone H3 and H4 acetylation and methylation mechanisms. Involved in several developmental processes including organization of plant organs, leaves formation, flowering time repression, and fertility. Modulates photoperiod-dependent epidermal leaves cell development; promotes cell division in long days, and cell expansion/division in short days. May be involved in the regulation of pathogenesis-related proteins (PRs).||[http://www.uniprot.org/uniprot/Q8LGE3 UniProtKB]||Am, V|| | + | | AT2G18915||ADAGIO 2, ADO2, LKP2, LOV KELCH PROTEIN 2||[http://arabidopsis.org/servlets/TairObject?id=500231567&type=locus TAIR]||Light Signaling||Scaffold00026 (length 499904) at 445582||0.0 |
| | |- | | |- |
| − | | ATBZIP27||Transcription factor||[http://www.genscript.com/cgi-bin/orf/refseq.pl?acc=NM_127331 GenScript]||Am, MI|| | + | | AT2G19520||ACG1, ATMSI4, FVE, MSI4, MULTICOPY SUPPRESSOR OF IRA1 4, NFC04, NFC4||[http://arabidopsis.org/servlets/TairObject?id=33019&type=locus TAIR]||Ambient Temperature, Autonomous||Scaffold00728 (length 157708) at 8716||6e-21 |
| | |- | | |- |
| − | | ATC||May form complexes with phosphorylated ligands by interfering with kinases and their effectors. Can substitute for TERMINAL FLOWER 1 (in vitro).||[http://www.uniprot.org/uniprot/Q9ZNV5 UniProtKB]|||| | + | | AT2G22540||AGAMOUS-LIKE 22, AGL22, SHORT VEGETATIVE PHASE, SVP||[http://arabidopsis.org/servlets/TairObject?id=31694&type=locus TAIR]||Ambient Temperature, Vernalization||Scaffold01187 (length 101153) at 47701||2e-24 |
| | |- | | |- |
| − | | ATCOL4||Sequence-specific DNA binding transcription factor activity, zinc ion binding.||[http://arabidopsis.org/servlets/TairObject?id=131285&type=locus TAIR]||L|| | + | | AT2G23380||CLF, CURLY LEAF, ICU1, INCURVATA 1, SDG1, SET1, SETDOMAIN 1, SETDOMAIN GROUP 1||[http://arabidopsis.org/servlets/TairObject?id=26534&type=locus TAIR]||Autonomous, Polycomb, Vernalization||Scaffold00354 (length 215005) at 64799||5e-41 |
| | |- | | |- |
| − | | ATCOL5||Sequence-specific DNA binding transcription factor activity, zinc ion binding.||[http://arabidopsis.org/servlets/TairObject?id=134422&type=locus TAIR]||L|| | + | | AT2G25930||EARLY FLOWERING 3, ELF3, PYK20||[http://arabidopsis.org/servlets/TairObject?id=32082&type=locus TAIR]||Light Signaling||Scaffold00371 (length 223748) at 73111||6e-19 |
| | |- | | |- |
| − | | ATSWC6||Encodes SERRATED LEAVES AND EARLY FLOWERING (SEF), an Arabidopsis homolog of the yeast SWC6 protein, a conserved subunit of the SWR1/SRCAP complex. SEF loss-of-function mutants have a pleiotropic phenotype characterized by serrated leaves, frequent absence of inflorescence internodes, bushy aspect, and flowers with altered number and size of organs. sef plants flower earlier than wild-type plants both under inductive and non-inductive photoperiods. SEF, ARP6 and PIE1 might form a molecular complex in Arabidopsis related to the SWR1/SRCAP complex identified in other eukaryotes.||[http://arabidopsis.org/servlets/TairObject?id=500231974&type=locus TAIR]||V|| | + | | AT2G32950||ARABIDOPSIS THALIANA CONSTITUTIVE PHOTOMORPHOGENIC 1, ATCOP1, CONSTITUTIVE PHOTOMORPHOGENIC 1, COP1, DEETIOLATED MUTANT 340, DET340, EMB168, EMBRYO DEFECTIVE 168, FUS1, FUSCA 1||[http://arabidopsis.org/servlets/TairObject?id=34462&type=locus TAIR]||Light Signaling||Scaffold00734 (length 158513) at 142779||5e-62 |
| | |- | | |- |
| − | | CAL||Probable transcription factor that promotes early floral meristem identity in synergy with APETALA1, FRUITFULL and LEAFY. Is required subsequently for the transition of an inflorescence meristem into a floral meristem. Seems to be partially redundant to the function of APETALA1.||[http://www.uniprot.org/uniprot/Q39375 UniProtKB]||FPI, MI|| | + | | AT2G33810||SPL3, SQUAMOSA PROMOTER BINDING PROTEIN-LIKE 3||[http://arabidopsis.org/servlets/TairObject?id=34194&type=locus TAIR]||Age||Scaffold00062 (length 451336) at 412489||2e-17 |
| | |- | | |- |
| − | | CCA1||Transcription factor involved in the circadian clock and in the phytochrome regulation. Binds to the promoter regions of APRR1/TOC1 and TCP21/CHE to repress their transcription. Binds to the promoter regions of CAB2A and CAB2B to promote their transcription. Represses both LHY and itself.||[http://www.uniprot.org/uniprot/P92973 UniProtKB]||L|| | + | | AT2G33835||FES1, FRIGIDA-ESSENTIAL 1||[http://arabidopsis.org/servlets/TairObject?id=500230872&type=locus TAIR]||Vernalization||Scaffold00102 (length 383343) at 88264||7e-17 |
| | |- | | |- |
| − | | CDF1||Cell growth defect factor||[http://www.uniprot.org/uniprot/Q4ACT9 UniProtKB]||L|| | + | | AT2G34140||||[http://arabidopsis.org/servlets/TairObject?id=33865&type=locus TAIR]||Light Signaling||Scaffold00270 (length 246371) at 215217||7e-49 |
| | |- | | |- |
| − | | CDF2||Cell growth defect factor||[http://www.uniprot.org/uniprot/Q3C1C7 UniProtKB]||L|| | + | | AT2G35670||FERTILIZATION INDEPENDENT SEED 2, FERTILIZATION-INDEPENDENT ENDOSPERM 2, FIE2, FIS2||[http://arabidopsis.org/servlets/TairObject?id=26543&type=locus TAIR]||Polycomb||Scaffold00857 (length 126644) at 55565||3e-04 |
| | |- | | |- |
| − | | CDF3||Plant non-specific lipid-transfer proteins transfer phospholipids as well as galactolipids across membranes. May play a role in wax or cutin deposition in the cell walls of expanding epidermal cells and certain secretory tissues.||[http://www.uniprot.org/uniprot/Q9S7I3 UniProtKB]||L|| | + | | AT2G40080||EARLY FLOWERING 4, ELF4||[http://arabidopsis.org/servlets/TairObject?id=34766&type=locus TAIR]||Light Signaling||Scaffold00254 (length 265810) at 228805||4e-24 |
| | |- | | |- |
| − | | CDF5||Cell growth defect factor||||L|| | + | | AT2G42200||ATSPL9, SPL9, SQUAMOSA PROMOTER BINDING PROTEIN-LIKE 9||[http://arabidopsis.org/servlets/TairObject?id=34557&type=locus TAIR]||Age||Scaffold00691 (length 141413) at 61795||5e-21 |
| | |- | | |- |
| − | | CHE||Transcription factor involved in the regulation of the circadian clock. Acts as a repressor of CCA1 by binding to its promoter. No binding to the LHY promoter.||[http://www.uniprot.org/uniprot/Q9FTA2 UniProtKB]||L|| | + | | AT2G43410||FPA||[http://arabidopsis.org/servlets/TairObject?id=34279&type=locus TAIR]||Autonomous||Scaffold01689 (length 75472) at 47663||4e-45 |
| | |- | | |- |
| − | | CIB1||Encodes a transcription factor CIB1 (cryptochrome-interacting basic-helix-loop-helix). CIB1 interacts with CRY2 (cryptochrome 2) in a blue light-specific manner in yeast and Arabidopsis cells, and it acts together with additional CIB1-related proteins to promote CRY2-dependent floral initiation. CIB1 positively regulates FT expression.||[http://arabidopsis.org/servlets/TairObject?id=130033&type=locus TAIR]||L|| | + | | AT2G46340||SPA1, SUPPRESSOR OF PHYA-105 1||[http://arabidopsis.org/servlets/TairObject?id=31336&type=locus TAIR]||Light Signaling||Scaffold00734 (length 158513) at 142779||3e-69 |
| | |- | | |- |
| − | | CKA1||Casein kinase II (CK2) catalytic subunit (alpha 1). One known substrate of CK2 is Phytochrome Interacting Factor 1 (PIF1). CK2-mediated phosphorylation enhances the light-induced degradation of PIF1 to promote photomorphogenesis.||[http://arabidopsis.org/servlets/TairObject?id=132478&type=locus TAIR]|||| | + | | AT2G46790||APRR9, ARABIDOPSIS PSEUDO-RESPONSE REGULATOR 9, PRR9, PSEUDO-RESPONSE REGULATOR 9, TL1, TOC1-LIKE PROTEIN 1||[http://arabidopsis.org/servlets/TairObject?id=32227&type=locus TAIR]||Light Signaling||Scaffold00001 (length 1030549) at 322801||1e-36 |
| | |- | | |- |
| − | | CKA2||Encodes the casein kinase II (CK2) catalytic subunit (alpha).||[http://arabidopsis.org/servlets/TairObject?id=37139&type=locus TAIR]|||| | + | | AT2G46830||ATCCA1, CCA1, CIRCADIAN CLOCK ASSOCIATED 1||[http://arabidopsis.org/servlets/TairObject?id=32221&type=locus TAIR]||Light Signaling||Scaffold00140 (length 354209) at 234299||7e-20 |
| | |- | | |- |
| − | | CLF||Similar to the product of the Polycomb-group gene Enhancer of zeste. Required for stable repression of AG and AP3. Putative role in cell fate determination. Involved in the control of leaf morphogenesis. mutants exhibit curled, involute leaves. AGAMOUS and APETALA3 are ectopically expressed in the mutant.||[http://arabidopsis.org/servlets/TairObject?id=26534&type=locus TAIR]||Au, V|| | + | | AT2G47700||RED AND FAR-RED INSENSITIVE 2, RFI2||[http://arabidopsis.org/servlets/TairObject?id=32070&type=locus TAIR]||Light Signaling||Scaffold01059 (length 101143) at 96557||1e-19 |
| | |- | | |- |
| − | | CO||Encodes a protein showing similarities to zinc finger transcription factors, involved in regulation of flowering under long days. Acts upstream of FT and SOC1.||[http://arabidopsis.org/servlets/TairObject?id=130492&type=locus TAIR]||FPI, L|| | + | | AT3G02380||ATCOL2, B-BOX DOMAIN PROTEIN 3, BBX3, COL2, CONSTANS-LIKE 2||[http://arabidopsis.org/servlets/TairObject?id=35635&type=locus TAIR]||Light Signaling||Scaffold01843 (length 51900) at 40767||3e-48 |
| | |- | | |- |
| − | | COL1||Homologous to the flowering-time gene CONSTANS.||[http://arabidopsis.org/servlets/TairObject?id=130495&type=locus TAIR]||FPI, L|| | + | | AT3G03450||RGA-LIKE 2, RGL2||[http://arabidopsis.org/servlets/TairObject?id=40021&type=locus TAIR]||Gibberellin||Scaffold00134 (length 346346) at 174722||0.0 |
| | |- | | |- |
| − | | COL2||Homologous to the flowering-time gene CONSTANS (CO) encoding zinc-finger proteins||[http://arabidopsis.org/servlets/TairObject?id=35635&type=locus TAIR]||FPI, L|| | + | | AT3G04610||FLK, FLOWERING LOCUS KH DOMAIN||[http://arabidopsis.org/servlets/TairObject?id=37455&type=locus TAIR]||Autonomous||Scaffold01384 (length 81068) at 28309||4e-29 |
| | |- | | |- |
| − | | COL3||Positive regulator of photomorphogenesis that acts downstream of COP1 but can promote lateral root development independently of COP1 and also function as a daylength-sensitive regulator of shoot branching.||[http://arabidopsis.org/servlets/TairObject?id=32704&type=locus TAIR]|||| | + | | AT3G05120||ATGID1A, GA INSENSITIVE DWARF1A, GID1A||[http://arabidopsis.org/servlets/TairObject?id=39457&type=locus TAIR]||Gibberellin||Scaffold00101 (length 425332) at 398762||4e-175 |
| | |- | | |- |
| − | | COL9||This gene belongs to the CO (CONSTANS) gene family. This gene family is divided in three subgroups: groups III, to which COL9 belongs, is characterised by one B-box (supposed to regulate protein-protein interactions) and a second diverged zinc finger. COL9 downregulates expression of CO (CONSTANS) as well as FT and SOC1 which are known regulatory targets of CO.||[http://arabidopsis.org/servlets/TairObject?id=38550&type=locus TAIR]||L|| | + | | AT3G07650||B-BOX DOMAIN PROTEIN 7, BBX7, COL9, CONSTANS-LIKE 9||[http://arabidopsis.org/servlets/TairObject?id=38550&type=locus TAIR]||Light Signaling||Scaffold00832 (length 123094) at 89599||1e-55 |
| | |- | | |- |
| − | | COP1||Represses photomorphogenesis and induces skotomorphogenesis in the dark. Contains a ring finger zinc-binding motif, a coiled-coil domain, and several WD-40 repeats, similar to G-beta proteins. The C-terminus has homology to TAFII80, a subunit of the TFIID component of the RNA polymerase II of Drosophila. Nuclear localization in the dark and cytoplasmic in the light.||[http://arabidopsis.org/servlets/TairObject?id=34462&type=locus TAIR]||L|| | + | | AT3G10390||FLD, FLOWERING LOCUS D||[http://arabidopsis.org/servlets/TairObject?id=35869&type=locus TAIR]||Ambient Temperature, Autonomous||Scaffold00232 (length 253418) at 139565||0.0 |
| | |- | | |- |
| − | | CRY1||Encodes CRY1, a flavin-type blue-light photoreceptor with ATP binding and autophosphorylation activity. Functions in perception of blue / green ratio of light. The photoreceptor may be involved in electron transport. Mutant phenotype displays a blue light-dependent inhibition of hypocotyl elongation. Photoreceptor activity requires light-induced homodimerisation of the N-terminal CNT1 domains of CRY1. Involved in blue-light induced stomatal opening. The C-terminal domain of the protein undergoes a light dependent conformational change. Also involved in response to circadian rhythm. Mutants exhibit long hypocotyl under blue light and are out of phase in their response to circadian rhythm. CRY1 is present in the nucleus and cytoplasm. Different subcellular pools of CRY1 have different functions during photomorphogenesis of Arabidopsis seedlings.||[http://arabidopsis.org/servlets/TairObject?id=129943&type=locus TAIR]||L||[http://www.uniprot.org/uniprot/B6REW9 Grape] | + | | AT3G12810||CHR13, PHOTOPERIOD-INDEPENDENT EARLY FLOWERING 1, PIE1, SRCAP||[http://arabidopsis.org/servlets/TairObject?id=37964&type=locus TAIR]||Vernalization||Scaffold00147 (length 345970) at 30164||0.0 |
| | |- | | |- |
| − | | CRY2||Blue light receptor mediating blue-light regulated cotyledon expansion and flowering time. Positive regulator of the flowering-time gene CONSTANS. This gene possesses a light-induced CNT2 N-terminal homodimerisation domain.Involved in blue-light induced stomatal opening. Involved in triggering chromatin decondensation. An 80-residue motif (NC80) is sufficient to confer CRY2's physiological function. It is proposed that the PHR domain and the C-terminal tail of the unphosphorylated CRY2 form a "closed" conformation to suppress the NC80 motif in the absence of light. In response to blue light, the C-terminal tail of CRY2 is phosphorylated and electrostatically repelled from the surface of the PHR domain to form an "open" conformation, resulting in derepression of the NC80 motif and signal transduction to trigger photomorphogenic responses. Cry2 phosphorylation and degradation both occur in the nucleus.||[http://arabidopsis.org/servlets/TairObject?id=28374&type=locus TAIR]||L|| | + | | AT3G15270||SPL5, SQUAMOSA PROMOTER BINDING PROTEIN-LIKE 5||[http://arabidopsis.org/servlets/TairObject?id=37842&type=locus TAIR]||Age||Scaffold00105 (length 392037) at 147284||5e-16 |
| | |- | | |- |
| − | | DPBF2||Basic leucine zipper transcription factor (BZIP67), identical to basic leucine zipper transcription factor GI:18656053 from (Arabidopsis thaliana); identical to cDNA basic leucine zipper transcription factor (atbzip67 gene) GI:18656052. Located in the nucleus and expressed during seed maturation in the cotyledons.||[http://arabidopsis.org/servlets/TairObject?id=35893&type=locus TAIR]|||| | + | | AT3G15354||SPA1-RELATED 3, SPA3||[http://arabidopsis.org/servlets/TairObject?id=500230681&type=locus TAIR]||Light Signaling||Scaffold00734 (length 158513) at 143450||1e-144 |
| | |- | | |- |
| − | | E12A11||Encodes a member of the FT and TFL1 family of phosphatidylethanolamine-binding proteins. It is expressed in seeds and up-regulated in response to ABA. Loss of function mutants show decreased rate of germination in the presence of ABA. ABA dependent regulation is mediated by both ABI3 and ABI5. ABI5 promotes MFT expression, primarily in the radicle-hypocotyl transition zone and ABI3 suppresses it in the seed.||[http://arabidopsis.org/servlets/TairObject?id=136229&type=locus TAIR]|||| | + | | AT3G18990||REDUCED VERNALIZATION RESPONSE 1, REM39, REPRODUCTIVE MERISTEM 39, VRN1||[http://arabidopsis.org/servlets/TairObject?id=37620&type=locus TAIR]||Vernalization||Scaffold00811 (length 108354) at 96661||2e-25 |
| | |- | | |- |
| − | | EEL||Transcription factor homologous to ABI5. Regulates AtEm1 expression by binding directly at the AtEm1 promoter. Located in the nucleus and expressed during seed maturation in the cotyledons and later in the whole embryo.||[http://arabidopsis.org/servlets/TairObject?id=35106&type=locus TAIR]|||| | + | | AT3G20740||FERTILIZATION-INDEPENDENT ENDOSPERM, FERTILIZATION-INDEPENDENT ENDOSPERM 1, FIE, FIE1, FIS3||[http://arabidopsis.org/servlets/TairObject?id=38691&type=locus TAIR]||Autonomous, Polycomb, Vernalization||Scaffold01670 (length 60304) at 9836||5e-20 |
| | |- | | |- |
| − | | EFS||Encodes a protein with histone lysine N-methyltransferase activity required specifically for the trimethylation of H3-K4 in FLC chromatin (and not in H3-K36 dimethylation). Acts as an inhibitor of flowering specifically involved in the autonomous promotion pathway. EFS also regulates the expression of genes involved in carotenoid biosynthesis.Modification of histone methylation at the CRTISO locus reduces transcript levels 90%. The increased shoot branching seen in some EFS mutants is likely due to the carotenoid biosynthesis defect having an effect on stringolactones.Required for ovule, embryo sac, anther and pollen development.||[http://arabidopsis.org/servlets/TairObject?id=136429&type=locus TAIR]||V|| | + | | AT3G21320||||[http://arabidopsis.org/servlets/TairObject?id=39170&type=locus TAIR]||Light Signaling||Scaffold00509 (length 192554) at 181887||1e-21 |
| | |- | | |- |
| − | | ELF3||Encodes a nuclear protein that is expressed rhythmically and interacts with phytochrome B to control plant development and flowering through a signal transduction pathway. Required component of the core circadian clock regardless of light conditions.||[http://arabidopsis.org/servlets/TairObject?id=32082&type=locus TAIR]||L|| | + | | AT3G24440||VERNALIZATION 5, VIL1, VIN3-LIKE 1, VRN5||[http://arabidopsis.org/servlets/TairObject?id=39196&type=locus TAIR]||Autonomous, Vernalization||Scaffold00396 (length 187979) at 73795||2e-69 |
| | |- | | |- |
| − | | ELF4||Encodes a novel nuclear 111 amino-acid phytochrome-regulated component of a negative feedback loop involving the circadian clock central oscillator components CCA1 and LHY. ELF4 is necessary for light-induced expression of both CCA1 and LHY, and conversely, CCA1 and LHY act negatively on light-induced ELF4 expression. ELF4 promotes clock accuracy and is required for sustained rhythms in the absence of daily light/dark cycles. It is involved in the phyB-mediated constant red light induced seedling de-etiolation process and may function to coregulate the expression of a subset of phyB-regulated genes.||[http://arabidopsis.org/servlets/TairObject?id=34766&type=locus TAIR]||L|| | + | | AT3G25730||EDF3, ETHYLENE RESPONSE DNA BINDING FACTOR 3||[http://arabidopsis.org/servlets/TairObject?id=37641&type=locus TAIR]||Light Signaling||Scaffold00930 (length 110378) at 63060||5e-100 |
| | |- | | |- |
| − | | ELF4-L3||Protein of unknown function||[http://arabidopsis.org/servlets/TairObject?id=32448&type=locus TAIR]|||| | + | | AT3G33520||ACTIN-RELATED PROTEIN 6, ARP6, ATARP6, EARLY IN SHORT DAYS 1, ESD1, SUF3, SUPPRESSOR OF FRI 3||[http://arabidopsis.org/servlets/TairObject?id=40458&type=locus TAIR]||Ambient Temperature, Vernalization||Scaffold00246 (length 286612) at 268521||1e-53 |
| | |- | | |- |
| − | | ELF4-L2||Protein of unknown function||[http://arabidopsis.org/servlets/TairObject?id=29911&type=locus TAIR]|||| | + | | AT3G46640||LUX, LUX ARRHYTHMO, PCL1, PHYTOCLOCK 1||[http://arabidopsis.org/servlets/TairObject?id=35747&type=locus TAIR]||Light Signaling||Scaffold01150 (length 88293) at 80272||3e-34 |
| | |- | | |- |
| − | | ELF4-L3||Protein of unknown function||[http://arabidopsis.org/servlets/TairObject?id=500231553&type=locus TAIR]||L|| | + | | AT3G47500||CDF3, CYCLING DOF FACTOR 3||[http://arabidopsis.org/servlets/TairObject?id=36439&type=locus TAIR]||Light Signaling||Scaffold00079 (length 471015) at 68175||1e-85 |
| | |- | | |- |
| − | | ELF4-L4||Protein of unknown function||[http://arabidopsis.org/servlets/TairObject?id=500231439&type=locus TAIR]|||| | + | | AT4G00650||FLA, FLOWERING LOCUS A, FRI, FRIGIDA||[http://arabidopsis.org/servlets/TairObject?id=128299&type=locus TAIR]||Vernalization||Scaffold00039 (length 505275) at 216632||4e-50 |
| | |- | | |- |
| − | | ELF5||Nuclear targeted protein involved in flowering time regulation that affects flowering time independent of FLC||[http://arabidopsis.org/servlets/TairObject?id=134377&type=locus TAIR]|||| | + | | AT4G02020||EZA1, SDG10, SET DOMAIN-CONTAINING PROTEIN 10, SWINGER, SWN||[http://arabidopsis.org/servlets/TairObject?id=129104&type=locus TAIR]||Autonomous, Polycomb, Vernalization||Scaffold00354 (length 215005) at 64799||2e-48 |
| | |- | | |- |
| − | | ELF6||Early Flowering 6 (ELF6) encodes a Jumonji N/C and zinc finger domain-containing protein that acts as a repressor in the photoperiod pathway. ELF6 interacts with BES1 in a Y2H assay, in vitro, and in Arabidosis protoplasts (based on BiFC). ELF6 may play a role in brassinosteroid signaling by affecting histone methylation in the promoters of BR-responsive genes.||[http://arabidopsis.org/servlets/TairObject?id=130928&type=locus TAIR]|||| | + | | AT4G02560||LD, LUMINIDEPENDENS||[http://arabidopsis.org/servlets/TairObject?id=26564&type=locus TAIR]||Autonomous||Scaffold00002 (length 840149) at 158601||4e-28 |
| | |- | | |- |
| − | | ELF9||Encodes a RNA binding protein ELF9 (EARLY FLOWERING9). Loss of ELF9 function in the Wassilewskija ecotype causes early flowering in short days. ELF9 reduces SOC1 (SUPPRESSOR OF OVEREXPRESSION OF CO1) transcript levels, possibly via nonsense-mediated mRNA decay.||[http://arabidopsis.org/servlets/TairObject?id=135649&type=locus TAIR]||FPI|| | + | | AT4G08920||ATCRY1, BLU1, BLUE LIGHT UNINHIBITED 1, CRY1, CRYPTOCHROME 1, ELONGATED HYPOCOTYL 4, HY4, OOP2, OUT OF PHASE 2||[http://arabidopsis.org/servlets/TairObject?id=129943&type=locus TAIR]||Light Signaling||Scaffold00331 (length 261439) at 124817||4e-128 |
| | |- | | |- |
| − | | EMF1||Involved in regulating reproductive development||[http://arabidopsis.org/servlets/TairObject?id=130620&type=locus TAIR]|||| | + | | AT4G11110||SPA1-RELATED 2, SPA2||[http://arabidopsis.org/servlets/TairObject?id=129633&type=locus TAIR]||Light Signaling||Scaffold01034 (length 107107) at 19205||2e-62 |
| | |- | | |- |
| − | | EMF2||Polycomb group protein with zinc finger domain involved in negative regulation of reproductive development. Forms a complex with FIE, CLF, and MSI1. This complex modulates the expression of target genes including AG, PI and AP3.||[http://arabidopsis.org/servlets/TairObject?id=134952&type=locus TAIR]|||| | + | | AT4G11880||AGAMOUS-LIKE 14, AGL14||[http://arabidopsis.org/servlets/TairObject?id=129748&type=locus TAIR]||Vernalization||Scaffold00249 (length 258199) at 131012||2e-29 |
| | |- | | |- |
| − | | ESD4||EARLY IN SHORT DAYS 4 Arabidopsis mutant shows extreme early flowering and alterations in shoot development. It encodes a SUMO protease, located predominantly at the periphery of the nucleus. Accelerates the transition from vegetative growth to flowering. Probably acts in the same pathway as NUA in affecting flowering time, vegetative and inflorescence development.||[http://arabidopsis.org/servlets/TairObject?id=128953&type=locus TAIR]|||| | + | | AT4G16250||PHYD, PHYTOCHROME D||[http://arabidopsis.org/servlets/TairObject?id=26567&type=locus TAIR]||Light Signaling||Scaffold00751 (length 152548) at 83707||0.0 |
| | |- | | |- |
| − | | FCA||Involved in the promotion of the transition of the vegetative meristem to reproductive development. Four forms of the protein (alpha, beta, delta and gamma) are produced by alternative splicing. Involved in RNA-mediated chromatin silencing. At one point it was believed to act as an abscisic acid receptor but the paper describing that function was retracted.||[http://arabidopsis.org/servlets/TairObject?id=128853&type=locus TAIR]||Am, Au||[http://www.uniprot.org/uniprot/D2Y3W8 Grape] | + | | AT4G16280||FCA||[http://arabidopsis.org/servlets/TairObject?id=128853&type=locus TAIR]||Ambient Temperature, Autonomous||Scaffold00104 (length 360147) at 120318||5e-13 |
| | |- | | |- |
| − | | FD||bZIP protein required for positive regulation of flowering. Mutants are late flowering. FD interacts with FT to promote flowering.Expressed in the shoot apex in floral anlagen, then declines in floral primordia.||[http://arabidopsis.org/servlets/TairObject?id=128068&type=locus TAIR]||Am, MI|| | + | | AT4G16845||REDUCED VERNALIZATION RESPONSE 2, VRN2||[http://arabidopsis.org/servlets/TairObject?id=228106&type=locus TAIR]||Autonomous, Polycomb, Vernalization||Scaffold00857 (length 126644) at 55203||3e-07 |
| | |- | | |- |
| − | | FES1||Encodes a zinc finger domain containing protein that is expressed in the shoot/root apex and vasculature, and acts with FRI to repress flowering.FES1 mutants in a Col(FRI+) background will flower early under inductive conditions.||[http://arabidopsis.org/servlets/TairObject?id=500230872&type=locus TAIR]||V|| | + | | AT4G18130||PHYE, PHYTOCHROME E||[http://arabidopsis.org/servlets/TairObject?id=26568&type=locus TAIR]||Light Signaling||Scaffold00751 (length 152548) at 83749||0.0 |
| | |- | | |- |
| − | | FIE1||Encodes a protein similar to the transcriptional regular of the animal Polycomb group and is involved in regulation of establishment of anterior-posterior polar axis in the endosperm and repression of flowering during vegetative phase. Mutation leads endosperm to develop in the absence of fertilization and flowers to form in seedlings and non-reproductive organs. Also exhibits maternal effect gametophytic lethal phenotype, which is suppressed by hypomethylation. Forms part of a large protein complex that can include VRN2 (VERNALIZATION 2), VIN3 (VERNALIZATION INSENSITIVE 3) and polycomb group proteins FERTILIZATION INDEPENDENT ENDOSPERM (FIE), CURLY LEAF (CLF) and SWINGER (SWN or EZA1). The complex has a role in establishing FLC (FLOWERING LOCUS C) repression during vernalization. In the ovule, the FIE transcript levels increase transiently just after fertilization.||[http://arabidopsis.org/servlets/TairObject?id=38691&type=locus TAIR]||Au, V|| | + | | AT4G20370||TSF, TWIN SISTER OF FT||[http://arabidopsis.org/servlets/TairObject?id=26553&type=locus TAIR]||Ambient Temperature||Scaffold00357 (length 260075) at 58807||1e-33 |
| | |- | | |- |
| − | | FIS2||Encodes a negative regulator of seed development in the absence of pollination. In the ovule, the FIS2 transcripts are accumulated at their highest level before fertilization and gradually decrease after fertilization.||[http://arabidopsis.org/servlets/TairObject?id=26543&type=locus TAIR]|||| | + | | AT4G22950||AGAMOUS-LIKE 19, AGL19, GL19||[http://arabidopsis.org/servlets/TairObject?id=128336&type=locus TAIR]||Vernalization||Scaffold00249 (length 258199) at 131012||4e-28 |
| | |- | | |- |
| − | | FKF1||Encodes FKF1, a flavin-binding kelch repeat F box protein, is clock-controlled, regulates transition to flowering. Forms a complex with GI on the CO promoter to regulate CO expression.||[http://arabidopsis.org/servlets/TairObject?id=137051&type=locus TAIR]||L|| | + | | AT4G24540||AGAMOUS-LIKE 24, AGL24||[http://arabidopsis.org/servlets/TairObject?id=127586&type=locus TAIR]||Vernalization||Scaffold01187 (length 101153) at 47701||2e-25 |
| | |- | | |- |
| − | | FLC||MADS-box protein encoded by FLOWERING LOCUS C - transcription factor that functions as a repressor of floral transition and contributes to temperature compensation of the circadian clock. Expression is downregulated during cold treatment. Vernalization, FRI and the autonomous pathway all influence the state of FLC chromatin. Both maternal and paternal alleles are reset by vernalization, but their earliest activation differs in timing and location. Histone H3 trimethylation at lysine 4 and histone acetylation are associated with active FLC expression, whereas histone deacetylation and histone H3 dimethylation at lysines 9 and 27 are involved in FLC repression. Expression is also repressed by two small RNAs (30- and 24-nt) complementary to the FLC sense strand 3’ to the polyA site. The small RNAs are most likely derived from an antisense transcript of FLC. Interacts with SOC1 and FT chromatin in vivo. Member of a protein complex.||[http://arabidopsis.org/servlets/TairObject?id=136002&type=locus TAIR]||Am, FPI, V||[http://www.uniprot.org/uniprot/D1MDP4 Grape] | + | | AT4G26000||PEP, PEPPER||[http://arabidopsis.org/servlets/TairObject?id=127403&type=locus TAIR]||Vernalization||Scaffold00021 (length 527983) at 157278||3e-29 |
| | |- | | |- |
| − | | FLD||Encodes a plant homolog of a SWIRM domain containing protein found in histone deacetylase complexes in mammals. Lesions in FLD result in hyperacetylation of histones in FLC chromatin, up-regulation of FLC expression and extremely delayed flowering. FLD plays a key role in regulating the reproductive competence of the shoot and results in different developmental phase transitions in Arabidopsis.||[http://arabidopsis.org/servlets/TairObject?id=35869&type=locus TAIR]||Am, Au|| | + | | AT4G30200||VEL1, VERNALIZATION5/VIN3-LIKE 1, VIL2, VIN3-LIKE 2||[http://arabidopsis.org/servlets/TairObject?id=128580&type=locus TAIR]||Autonomous, Vernalization||Scaffold00026 (length 499904) at 369417||7e-64 |
| | |- | | |- |
| − | | FLK||RNA binding, nucleic acid binding; positive regulation of flower development||[http://arabidopsis.org/servlets/TairObject?id=37455&type=locus TAIR]||Au|| | + | | AT4G34530||CIB1, CRYPTOCHROME-INTERACTING BASIC-HELIX-LOOP-HELIX 1||[http://arabidopsis.org/servlets/TairObject?id=130033&type=locus TAIR]||Light Signaling||Scaffold01322 (length 99420) at 82955||2e-29 |
| | |- | | |- |
| − | | FLP1*||Encodes a small protein with unknown function and is similar to flower promoting factor 1. This gene is not expressed in apical meristem after floral induction but is expressed in roots, flowers, and in low abundance, leaves.||[http://arabidopsis.org/servlets/TairObject?id=128483&type=locus TAIR]|||| | + | | AT4G35900||ATBZIP14, FD, FD-1||[http://arabidopsis.org/servlets/TairObject?id=128068&type=locus TAIR]||Ambient Temperature||Scaffold00367 (length 240396) at 113139||1e-16 |
| | |- | | |- |
| − | | FPA||FPA is a gene that regulates flowering time in Arabidopsis via a pathway that is independent of daylength (the autonomous pathway). Mutations in FPA result in extremely delayed flowering. Double mutants with FCA have reduced fertility and single/double mutants have defects in siRNA mediated chromatin silencing.||[http://arabidopsis.org/servlets/TairObject?id=34279&type=locus TAIR]||Au|| | + | | AT5G02810||APRR7, PRR7, PSEUDO-RESPONSE REGULATOR 7||[http://arabidopsis.org/servlets/TairObject?id=131538&type=locus TAIR]||Light Signaling||Scaffold00125 (length 356885) at 276518||1e-33 |
| | |- | | |- |
| − | | FPF1||Encodes a small protein of 12.6 kDa that regulates flowering and is involved in gibberellin signalling pathway. It is expressed in apical meristems immediately after the photoperiodic induction of flowering. Genetic interactions with flowering time and floral organ identity genes suggest that this gene may be involved in modulating the competence to flower. There are two other genes similar to FPF1, FLP1 (At4g31380) and FLP2 (no locus name yet, on BAC F8F16 on chr 4). This is so far a plant-specific gene and is only found in long-day mustard, arabidopsis, and rice.||[http://arabidopsis.org/servlets/TairObject?id=131288&type=locus TAIR]|||| | + | | AT5G03840||TERMINAL FLOWER 1, TFL1||[http://arabidopsis.org/servlets/TairObject?id=131459&type=locus TAIR]||Ambient Temperature||Scaffold00181 (length 337602) at 12825||1e-28 |
| | |- | | |- |
| − | | FRI||Encodes a major determinant of natural variation in Arabidopsis flowering time. Dominant alleles of FRI confer a vernalization requirement causing plants to overwinter vegetatively. Many early flowering accessions carry loss-of-function fri alleles .Twenty distinct haplotypes that contain non-functional FRI alleles have been identified and the distribution analyzed in over 190 accessions. The common lab strains- Col and Ler each carry loss of function mutations in FRI.||[http://arabidopsis.org/servlets/TairObject?id=128299&type=locus TAIR]||V|| | + | | AT5G08330||CCA1 HIKING EXPEDITION, CHE, TRANSCRIPTION FACTOR TCP21, TCP21||[http://www.uniprot.org/uniprot/Q9FTA2 UniProtKB]||Light Signaling||Scaffold00993 (length 109486) at 294||5e-27 |
| | |- | | |- |
| − | | FRL1||Encodes a sterol-C24-methyltransferases involved in sterol biosynthesis. Mutants display altered sterol composition, serrated petals and sepals and altered cotyledon vascular patterning as well as ectopic endoreduplication. This suggests that suppression of endoreduplication is important for petal morphogenesis and that normal sterol composition is required for this suppression.||[http://arabidopsis.org/servlets/TairObject?id=27665&type=locus TAIR]||V|| | + | | AT5G10140||AGAMOUS-LIKE 25, AGL25, FLC, FLF, FLOWERING LOCUS C, FLOWERING LOCUS F||[http://arabidopsis.org/servlets/TairObject?id=136002&type=locus TAIR]||Ambient Temperature, Vernalization||Scaffold10765 (length 2447) at 708||8e-22 |
| | |- | | |- |
| − | | FRL2||Family member of FRI-related genes that is required for the winter-annual habit. ||[http://arabidopsis.org/servlets/TairObject?id=226970&type=locus TAIR]||V|| | + | | AT5G11530||EMBRYONIC FLOWER 1, EMF1||[http://arabidopsis.org/servlets/TairObject?id=130620&type=locus TAIR]||Polycomb||Scaffold00253 (length 240879) at 11069||1e-06 |
| | |- | | |- |
| − | | FT||FT, together with LFY, promotes flowering and is antagonistic with its homologous gene, TERMINAL FLOWER1 (TFL1). FT is expressed in leaves and is induced by long day treatment. Either the FT mRNA or protein is translocated to the shoot apex where it induces its own expression. Recent data suggests that FT protein acts as a long-range signal. FT is a target of CO and acts upstream of SOC1.||[http://arabidopsis.org/servlets/TairObject?id=30541&type=locus TAIR]||Am, FPI||[http://www.uniprot.org/uniprot/D1MDP3 Grape], [http://www.uniprot.org/uniprot/E5BBS7 Strawberry] | + | | AT5G13480||FY||[http://arabidopsis.org/servlets/TairObject?id=136108&type=locus TAIR]||Autonomous||Scaffold00166 (length 328538) at 74796||2e-43 |
| | |- | | |- |
| − | | FUL||MADS box gene negatively regulated by APETALA1||[http://arabidopsis.org/servlets/TairObject?id=134558&type=locus TAIR]||MI||[http://www.uniprot.org/uniprot/D1MDP9 Grape] | + | | AT5G15840||B-BOX DOMAIN PROTEIN 1, BBX1, CO, CONSTANS, FG||[http://arabidopsis.org/servlets/TairObject?id=130492&type=locus TAIR]||Light Signaling||Scaffold01843 (length 51900) at 40767||3e-29 |
| | |- | | |- |
| − | | FVE||Controls flowering.||[http://arabidopsis.org/servlets/TairObject?id=33019&type=locus TAIR]||Am, Au|| | + | | AT5G15850||ATCOL1, B-BOX DOMAIN PROTEIN 2, BBX2, COL1, CONSTANS-LIKE 1||[http://arabidopsis.org/servlets/TairObject?id=130495&type=locus TAIR]||Light Signaling||Scaffold01843 (length 51900) at 40767||5e-44 |
| | |- | | |- |
| − | | FY||Encodes a protein with similarity to yeast Pfs2p, an mRNA processing factor. Involved in regulation of flowering time; affects FCA mRNA processing. Homozygous mutants are late flowering, null alleles are embryo lethal.||[http://arabidopsis.org/servlets/TairObject?id=136108&type=locus TAIR]||Au|| | + | | AT5G17690||ATLHP1, LHP1, LIKE HETEROCHROMATIN PROTEIN 1, TERMINAL FLOWER 2, TFL2||[http://arabidopsis.org/servlets/TairObject?id=134923&type=locus TAIR]||Polycomb||Scaffold00696 (length 140617) at 128983||7e-13 |
| | |- | | |- |
| − | | GBF4||Encodes a basic leucine zipper G-box binding factor that can bind to G-box motifs only as heterodimers with GBF2 or GBF3. A single amino acid change can confer G-box binding as homodimers.||[http://arabidopsis.org/servlets/TairObject?id=28911&type=locus TAIR]|||| | + | | AT5G23150||ENHANCER OF AG-4 2, HUA2||[http://arabidopsis.org/servlets/TairObject?id=135220&type=locus TAIR]||Vernalization||Scaffold00686 (length 145647) at 102689||5e-42 |
| | |- | | |- |
| − | | GI||Together with CONSTANTS (CO) and FLOWERING LOCUS T (FT), GIGANTEA promotes flowering under long days in a circadian clock-controlled flowering pathway. GI acts earlier than CO and FT in the pathway by increasing CO and FT mRNA abundance. Located in the nucleus. Regulates several developmental processes, including photoperiod-mediated flowering, phytochrome B signaling, circadian clock, carbohydrate metabolism, and cold stress response. The gene's transcription is controlled by the circadian clock and it is post-transcriptionally regulated by light and dark. Forms a complex with FKF1 on the CO promoter to regulate CO expression.||[http://arabidopsis.org/servlets/TairObject?id=136959&type=locus TAIR]||L|| | + | | AT5G24470||APRR5, PRR5, PSEUDO-RESPONSE REGULATOR 5||[http://arabidopsis.org/servlets/TairObject?id=135985&type=locus TAIR]||Light Signaling||Scaffold00001 (length 1030549) at 322798||3e-40 |
| | |- | | |- |
| − | | GRF1||Growth regulating factor encoding transcription activator. One of the nine members of a GRF gene family, containing nuclear targeting domain. Mutants result in smaller leaves indicating the role of the gene in leaf development. Expressed in root, shoot and flower||[http://arabidopsis.org/servlets/TairObject?id=34438&type=locus TAIR]||FPI|| | + | | AT5G24930||ATCOL4, B-BOX DOMAIN PROTEIN 5, BBX5, COL4, CONSTANS-LIKE 4||[http://arabidopsis.org/servlets/TairObject?id=131285&type=locus TAIR]||Light Signaling||Scaffold11225 (length 3326) at 2816||8e-23 |
| | |- | | |- |
| − | | GRF10*||Encodes a 14-3-3 protein. This protein is reported to interact with the BZR1 transcription factor involved in brassinosteroid signaling and may affect the nucleocytoplasmic shuttling of BZR1||[http://arabidopsis.org/servlets/TairObject?id=136501&type=locus TAIR]||FPI|| | + | | AT5G35840||PHYC, PHYTOCHROME C||[http://arabidopsis.org/servlets/TairObject?id=133466&type=locus TAIR]||Light Signaling||Scaffold01070 (length 102706) at 75395||0.0 |
| | |- | | |- |
| − | | GRF11*||Encodes a 14-3-3 protein. Binds H+-ATPase in response to blue light.||[http://arabidopsis.org/servlets/TairObject?id=26877&type=locus TAIR]||FPI|| | + | | AT5G37055||ATSWC6, SEF, SERRATED LEAVES AND EARLY FLOWERING||[http://arabidopsis.org/servlets/TairObject?id=500231974&type=locus TAIR]||Vernalization||Scaffold00925 (length 141061) at 38143||4e-29 |
| | |- | | |- |
| − | | GRF12*||Encodes a 14-3-3 protein.||[http://arabidopsis.org/servlets/TairObject?id=136716&type=locus TAIR]||FPI|| | + | | AT5G39660||CDF2, CYCLING DOF FACTOR 2||[http://arabidopsis.org/servlets/TairObject?id=133414&type=locus TAIR]||Light Signaling||Scaffold00651 (length 145047) at 19066||1e-90 |
| | |- | | |- |
| − | | GRF2||G-box binding factor GF14 omega encoding a 14-3-3 protein||[http://arabidopsis.org/servlets/TairObject?id=30202&type=locus TAIR]||FPI|| | + | | AT5G42790||ARS5, ARSENIC TOLERANCE 5, ATPSM30, PAF1, PROTEASOME ALPHA SUBUNIT F1||[http://arabidopsis.org/servlets/TairObject?id=133505&type=locus TAIR]||Vernalization||Scaffold00528 (length 197283) at 110933||5e-64 |
| | |- | | |- |
| − | | GRF3||Growth regulating factor encoding transcription activator. One of the nine members of a GRF gene family, containing nuclear targeting domain. Mutants result in smaller leaves indicating the role of the gene in leaf development. Expressed in root, shoot and flower.||[http://arabidopsis.org/servlets/TairObject?id=32311&type=locus TAIR]||FPI|| | + | | AT5G51230||ATEMF2, CYR1, CYTOKININ RESISTANT 1, EMBRYONIC FLOWER 2, EMF2, VEF2||[http://arabidopsis.org/servlets/TairObject?id=134952&type=locus TAIR]||Polycomb||Scaffold00857 (length 126644) at 44887||9e-19 |
| | |- | | |- |
| − | | GRF4||GF14 protein phi chain member of 14-3-3 protein family. This protein is reported to interact with the BZR1 transcription factor involved in brassinosteroid signaling and may affect the nucleocytoplasmic shuttling of BZR1||[http://arabidopsis.org/servlets/TairObject?id=137475&type=locus TAIR]||FPI|| | + | | AT5G57360||ADAGIO 1, ADO1, FKF1-LIKE PROTEIN 2, FKL2, LKP1, LOV KELCH PROTEIN 1, ZEITLUPE, ZTL||[http://arabidopsis.org/servlets/TairObject?id=134481&type=locus TAIR]||Light Signaling||Scaffold00026 (length 499904) at 445597||0.0 |
| | |- | | |- |
| − | | GRF5||Growth regulating factor encoding transcription activator. One of the nine members of a GRF gene family, containing nuclear targeting domain. Involved in leaf development and expressed in root, shoot and flower.||[http://arabidopsis.org/servlets/TairObject?id=38052&type=locus TAIR]||FPI|| | + | | AT5G57380||VERNALIZATION INSENSITIVE 3, VIN3||[http://arabidopsis.org/servlets/TairObject?id=134483&type=locus TAIR]||Autonomous, Vernalization||Scaffold00026 (length 499904) at 369405||3e-48 |
| | |- | | |- |
| − | | GRF6||Growth regulating factor encoding transcription activator. One of the nine members of a GRF gene family, containing nuclear targeting domain. Involved in leaf development and expressed in root, shoot and flower.||[http://arabidopsis.org/servlets/TairObject?id=33231&type=locus TAIR]||FPI|| | + | | AT5G57660||ATCOL5, B-BOX DOMAIN PROTEIN 6, BBX6, COL5, CONSTANS-LIKE 5||[http://arabidopsis.org/servlets/TairObject?id=134422&type=locus TAIR]||Light Signaling||Scaffold11225 (length 3326) at 2816||1e-25 |
| | |- | | |- |
| − | | GRF7||Encodes GF14 ν, a 14-3-3 protein isoform (14-3-3ν).||[http://arabidopsis.org/servlets/TairObject?id=36055&type=locus TAIR]||FPI|| | + | | AT5G58230||ARABIDOPSIS MULTICOPY SUPRESSOR OF IRA1, ATMSI1, MATERNAL EFFECT EMBRYO ARREST 70, MEE70, MSI1, MULTICOPY SUPRESSOR OF IRA1||[http://arabidopsis.org/servlets/TairObject?id=132921&type=locus TAIR]||Autonomous, Polycomb, Vernalization||Scaffold00615 (length 179658) at 161081||2e-61 |
| | |- | | |- |
| − | | GRF8||Growth regulating factor encoding transcription activator. One of the nine members of a GRF gene family, containing nuclear targeting domain. Involved in leaf development and expressed in shoot and flower.||[http://arabidopsis.org/servlets/TairObject?id=129479&type=locus TAIR]||FPI|| | + | | AT5G60100||APRR3, PRR3, PSEUDO-RESPONSE REGULATOR 3||[http://arabidopsis.org/servlets/TairObject?id=133317&type=locus TAIR]||Light Signaling||Scaffold02075 (length 43325) at 14139||3e-25 |
| | |- | | |- |
| − | | HAP3A||Encodes a transcription factor from the nuclear factor Y (NF-Y) family, AtNF-YB1. Confers drought tolerance.||[http://arabidopsis.org/servlets/TairObject?id=35360&type=locus TAIR]||FPI|| | + | | AT5G61380||APRR1, ATTOC1, PRR1, PSEUDO-RESPONSE REGULATOR 1, TIMING OF CAB EXPRESSION 1, TOC1||[http://arabidopsis.org/servlets/TairObject?id=133196&type=locus TAIR]||Light Signaling||Scaffold00753 (length 123742) at 70468||2e-47 |
| | |- | | |- |
| − | | HAP3B||||||FPI|| | + | | AT5G62430||CDF1, CYCLING DOF FACTOR 1||[http://arabidopsis.org/servlets/TairObject?id=131911&type=locus TAIR]||Light Signaling||Scaffold01102 (length 99830) at 51435||1e-59 |
| | |- | | |- |
| − | | HAP5A||Encodes a protein with similarity to a subunit of the CCAAT promoter motif binding complex of yeast.One of two members of this class (HAP5A) and expressed in vegetative and reproductive tissues.||[http://arabidopsis.org/servlets/TairObject?id=126547&type=locus TAIR]||FPI|| | + | | AT5G65050||AGAMOUS-LIKE 31, AGL31, MADS AFFECTING FLOWERING 2, MAF2||[http://arabidopsis.org/servlets/TairObject?id=135154&type=locus TAIR]||Ambient Temperature, Vernalization||Scaffold10765 (length 2447) at 723||4e-20 |
| | |- | | |- |
| − | | HAP5B||Encodes a protein with similarity to a subunit of the CCAAT promoter motif binding complex of yeast.One of two members of this class (HAP5B) and expressed in vegetative and reproductive tissues||[http://arabidopsis.org/servlets/TairObject?id=27444&type=locus TAIR]||FPI|| | + | | AT5G65060||AGAMOUS-LIKE 70, AGL70, FCL3, MADS AFFECTING FLOWERING 3, MAF3||[http://arabidopsis.org/servlets/TairObject?id=130633&type=locus TAIR]||Ambient Temperature, Vernalization||Scaffold10765 (length 2447) at 639||7e-15 |
| | |- | | |- |
| − | | HAP5C||Heme activated protein (HAP5c)||[http://arabidopsis.org/servlets/TairObject?id=30861&type=locus TAIR]||FPI|| | + | | AT5G65070||AGAMOUS-LIKE 69, AGL69, FCL4, MADS AFFECTING FLOWERING 4, MAF4||[http://arabidopsis.org/servlets/TairObject?id=130634&type=locus TAIR]||Ambient Temperature, Vernalization||Scaffold10765 (length 2447) at 723||6e-20 |
| | |- | | |- |
| − | | HUA2||Putative transcription factor. Member of the floral homeotic AGAMOUS pathway.Mutations in HUA enhance the phenotype of mild ag-4 allele. Single hua mutants are early flowering and have reduced levels of FLC mRNA. Other MADS box flowering time genes such as FLM and MAF2 also appear to be regulated by HUA2. HUA2 normally activates FLC expression and enhances AG function.||[http://arabidopsis.org/servlets/TairObject?id=135220&type=locus TAIR]||V|| | + | | AT5G65080||AGAMOUS-LIKE 68, AGL68, MADS AFFECTING FLOWERING 5, MAF5||[http://arabidopsis.org/servlets/TairObject?id=130635&type=locus TAIR]||Ambient Temperature, Vernalization||Scaffold10765 (length 2447) at 699||1e-18 |
| − | |-
| |
| − | | LD||Encodes a nuclear localized protein with similarity to transcriptional regulators. Recessive mutants are late flowering. Expression of LFY is reduced in LD mutants.||[http://arabidopsis.org/servlets/TairObject?id=26564&type=locus TAIR]||Au||
| |
| − | |-
| |
| − | | LDL1*||Encodes a homolog of human Lysine-Specific Demethylase1. Involved in H3K4 methylation of target genes including the flowering time loci FLC and FWA. Located in nucleus. Negatively regulates root elongation. Involved in repression of LRP1 via histone deacetylation.||[http://arabidopsis.org/servlets/TairObject?id=29274&type=locus TAIR]||Am, Au||
| |
| − | |-
| |
| − | | LDL2*||Encodes a homolog of human Lysine-Specific Demethylase1. Involved in H3K4 methylation of target genes including the flowering loci FLC and FWA.||[http://arabidopsis.org/servlets/TairObject?id=38624 TAIR]||Am, Au||
| |
| − | |-
| |
| − | | LFY||Encodes transcriptional regulator that promotes the transition to flowering.Involved in floral meristem development. LFY is involved in the regulation of AP3 expression, and appears to bring the F-box protein UFO to the AP3 promoter. Amino acids 46-120 define a protein domain that mediates self-interaction.||[http://arabidopsis.org/servlets/TairObject?id=132616&type=locus TAIR]||FPI, MI||[http://www.uniprot.org/uniprot/Q8L5W6 Grape], [http://www.uniprot.org/uniprot/C4NF85 Strawberry]
| |
| − | |-
| |
| − | | LHP1||Regulates the meristem response to light signals and the maintenance of inflorescence meristem identity. Influences developmental processes controlled by APETALA1. TFL2 silences specific genes within euchromatin but not genes positioned in heterochromatin. TFL2 protein localized preferentially to euchromatic regions and not to heterochromatic chromocenters. Involved in euchromatin organization. Required for epigenetic maintenance of the vernalized state.||[http://arabidopsis.org/servlets/TairObject?id=134923&type=locus TAIR]||||
| |
| − | |-
| |
| − | | LHY||LHY encodes a myb-related putative transcription factor involved in circadian rhythm along with another myb transcription factor CCA1.||[http://arabidopsis.org/servlets/TairObject?id=137165&type=locus TAIR]||L||
| |
| − | |-
| |
| − | | LKP2||Encodes a member of F-box proteins that includes two other proteins in Arabidopsis (ZTL and FKF1). These proteins contain a unique structure containing a PAS domain at their N-terminus, an F-box motif, and 6 kelch repeats at their C-terminus. Overexpression results in arrhythmic phenotypes for a number of circadian clock outputs in both constant light and constant darkness, long hypocotyls under multiple fluences of both red and blue light, and a loss of photoperiodic control of flowering time. Although this the expression of this gene itself is not regulated by circadian clock, it physically interacts with Dof transcription factors that are transcriptionally regulated by circadian rhythm. LKP2 interacts with Di19, CO/COL family proteins.||[http://arabidopsis.org/servlets/TairObject?id=500231567&type=locus TAIR]||L||
| |
| − | |-
| |
| − | | LUX||Encodes a myb family transcription factor with a single Myb DNA-binding domain (type SHAQKYF) that is unique to plants and is essential for circadian rhythms, specifically for transcriptional regulation within the circadian clock. LUX is required for normal rhythmic expression of multiple clock outputs in both constant light and darkness. It is coregulated with TOC1 and seems to be repressed by CCA1 and LHY by direct binding of these proteins to the evening element in the LUX promoter.||[http://arabidopsis.org/servlets/TairObject?id=35747&type=locus TAIR]||L||
| |
| − | |-
| |
| − | | MAF1||MADS domain protein - flowering regulator that is closely related to FLC. Deletion of this locus in Nd ecotype is correlated with earlier flowering in short days suggesting function as a negative regulator of flowering.||[http://arabidopsis.org/servlets/TairObject?id=29106&type=locus TAIR]||Am, FPI, V||
| |
| − | |-
| |
| − | | MAF3||MADS domain protein - flowering regulator that is closely related to FLC||[http://arabidopsis.org/servlets/TairObject?id=130633&type=locus TAIR]||Am, FPI, V||
| |
| − | |-
| |
| − | | MAF4||Encodes MADS-box containing FLC paralog. Five splice variants have been identified but not characterized with respect to expression patterns and/or differing function. Overexpression of the gene in the Landsberg ecotype leads to a delay in flowering, transcript levels of MAF4 are reduced after a 6 week vernalization.||[http://arabidopsis.org/servlets/TairObject?id=130634&type=locus TAIR]||Am, FPI, V||
| |
| − | |-
| |
| − | | MAF5||Is upregulated during vernalization and regulates flowering time. Encodes MADS-domain protein. Two variants encoding proteins of 198 and 184 amino acids have been reported.||[http://arabidopsis.org/servlets/TairObject?id=130635&type=locus TAIR]||Am, FPI, V||
| |
| − | |-
| |
| − | | MEA*||Encodes a putative transcription factor MEDEA (MEA) that negatively regulates seed development in the absence of fertilization. Mutations in this locus result in embryo lethality. MEA is a Polycomb group gene that is imprinted in the endosperm. The maternal allele is expressed and the paternal allele is silent. MEA is controlled by DEMETER (DME), a DNA glycosylase required to activate MEA expression, and METHYLTRANSFERASE I (MET1), which maintains CG methylation at the MEA locus. MEA is involved in the negative regulation of its own imprinted gene expression; the effect is not only allele-specific but also dynamically regulated during seed development. In the ovule, the MEA transcripts are accumulated at their highest level before fertilization and gradually decrease after fertilization||[http://arabidopsis.org/servlets/TairObject?id=136450&type=locus TAIR]||Au, V||
| |
| − | |-
| |
| − | | MSI1||Encodes a WD-40 repeat containing protein that functions in chromatin assembly as part of the CAF1 and FIE complex. Mutants exhibit parthenogenetic development that includes proliferation of unfertilized endosperm and embryos. In heterozygous plants 50% of embryos abort. Of the aborted embryos the early aborted class are homozygous and the later aborting lass are heterozygotes in which the defective allele is maternally transmitted. Other phenotypes include defects in ovule morphogenesis and organ initiation,as well as increased levels of heterochromatic DNA. MSI1 is needed for the transition to flowering. In Arabidopsis, the three CAF-1 subunits are encoded by FAS1, FAS2 and, most likely, MSI1, respectively. Mutations in FAS1 or FAS2 lead to increased frequency of homologous recombination and T-DNA integration in Arabidopsis. In the ovule, the MSI1 transcripts are accumulated at their highest level before fertilization and gradually decrease after fertilization. MSI is biallelically expressed, the paternall allele is expressed in the endosperm and embryo and is not imprinted. MSI1 forms a complex with RBR1 that is required for activation of the imprinted genes FIS2 and FWA. This activation is mediated by MSI1/RBR1 mediated repression of MET1.||[http://arabidopsis.org/servlets/TairObject?id=132921&type=locus TAIR]||Au, V||
| |
| − | |-
| |
| − | | MSI2*||Core histone-binding subunit that may target chromatin assembly factors, chromatin remodeling factors and histone deacetylases to their histone substrates in a manner that is regulated by nucleosomal DNA||[http://www.uniprot.org/uniprot/O22468 UniProtKB]||Au, V||
| |
| − | |-
| |
| − | | MSI3*||Core histone-binding subunit that may target chromatin assembly factors, chromatin remodeling factors and histone deacetylases to their histone substrates in a manner that is regulated by nucleosomal DNA||[http://www.uniprot.org/uniprot/O22469 UniProtKB]||Au, V||
| |
| − | |-
| |
| − | | NF-YB10||Sequence-specific DNA binding transcription factor activity; regulation of transcription, DNA-dependent||[http://arabidopsis.org/servlets/TairObject?id=37281&type=locus TAIR]||FPI||
| |
| − | |-
| |
| − | | NF-YB3||Sequence-specific DNA binding transcription factor activity; regulation of transcription, DNA-dependent||[http://arabidopsis.org/servlets/TairObject?id=128758&type=locus TAIR]||FPI||
| |
| − | |-
| |
| − | | NF-YB7||Sequence-specific DNA binding transcription factor activity; regulation of transcription, DNA-dependent||[http://arabidopsis.org/servlets/TairObject?id=33626&type=locus TAIR]||FPI||
| |
| − | |-
| |
| − | | NF-YB8||Sequence-specific DNA binding transcription factor activity; regulation of transcription, DNA-dependent||[http://arabidopsis.org/servlets/TairObject?id=34862&type=locus TAIR]||FPI||
| |
| − | |-
| |
| − | | NF-YC3||Sequence-specific DNA binding transcription factor activity; regulation of transcription, DNA-dependent||[http://arabidopsis.org/servlets/TairObject?id=27340&type=locus TAIR]||FPI||
| |
| − | |-
| |
| − | | NF-YC4||Sequence-specific DNA binding transcription factor activity; regulation of transcription, DNA-dependent||[http://arabidopsis.org/servlets/TairObject?id=133713&type=locus TAIR]||FPI||
| |
| − | |-
| |
| − | | NFC5*||Cell cycle-related repressor genes encoding WD-repeat proteins.||[http://arabidopsis.org/servlets/TairObject?id=129407&type=locus TAIR]||Am, Au||
| |
| − | |-
| |
| − | | PAF1||Encodes a protein with extensive homology to the largest subunit of the multicatalytic proteinase complex (proteasome). Negatively regulates thiol biosynthesis and arsenic tolerance.||[http://arabidopsis.org/servlets/TairObject?id=133505&type=locus TAIR]||V||
| |
| − | |-
| |
| − | | PAF2||Encodes 20S proteasome subunit PAF2 (PAF2).||[http://arabidopsis.org/servlets/TairObject?id=30983&type=locus TAIR]||V||
| |
| − | |-
| |
| − | | PEP||Encodes a novel Arabidopsis gene encoding a polypeptide with K-homology (KH) RNA-binding modules, which acts on vegetative growth and pistil development. Genetic studies suggest that PEP interacts with element(s) of the CLAVATA signaling pathway.||[http://arabidopsis.org/servlets/TairObject?id=127403&type=locus TAIR]||V||
| |
| − | |-
| |
| − | | PFT1||Encodes a nuclear protein that acts in a phyB pathway (but downstream of phyB) and induces flowering in response to suboptimal light conditions. PFT1 promotes flowering in CO dependent and independent pathways and integrates several environmental stimuli, such as light quality and JA-dependent defenses. Mutants are hypo-responsive to far-red and hyper-responsive to red light and flower late under long day conditions. Also shown to be a Mediator subunit regulating jasmonate-dependent defense.||[http://arabidopsis.org/servlets/TairObject?id=30064&type=locus TAIR]||||
| |
| − | |-
| |
| − | | PHYA||Light-labile cytoplasmic red/far-red light photoreceptor involved in the regulation of photomorphogenesis. It exists in two inter-convertible forms: Pr and Pfr (active) and functions as a dimer.The N terminus carries a single tetrapyrrole chromophore, and the C terminus is involved in dimerization. It is the sole photoreceptor mediating the FR high irradiance response (HIR). Major regulator in red-light induction of phototropic enhancement. Involved in the regulation of de-etiolation. Involved in gravitropism and phototropism. Requires FHY1 for nuclear accumulation.||[http://arabidopsis.org/servlets/TairObject?id=27545&type=locus TAIR]||L||[http://www.uniprot.org/uniprot/A5B9C2 Grape]
| |
| − | |-
| |
| − | | PHYB||Red/far-red photoreceptor involved in the regulation of de-etiolation. Exists in two inter-convertible forms: Pr and Pfr (active). Involved in the light-promotion of seed germination and in the shade avoidance response. Promotes seedling etiolation in both the presence and absence of phytochrome A. Overexpression results in etiolation under far-red light. Accumulates in the nucleus after exposure to far red light.||[http://arabidopsis.org/servlets/TairObject?id=26548&type=locus TAIR]||L||[http://www.uniprot.org/uniprot/B9U4G3 Grape]
| |
| − | |-
| |
| − | | PHYC||Encodes the apoprotein of phytochrome;one of a family of photoreceptors that modulate plant growth and development.||[http://arabidopsis.org/servlets/TairObject?id=133466&type=locus TAIR]||L||[http://www.uniprot.org/uniprot/B9U4G4 Grape]
| |
| − | |-
| |
| − | | PHYD||Encodes a phytochrome photoreceptor with a function similar to that of phyB that absorbs the red/far-red part of the light spectrum and is involved in light responses. It cannot compensate for phyB loss in Arabidopsis but can substitute for tobacco phyB in vivo.||[http://arabidopsis.org/servlets/TairObject?id=26567&type=locus TAIR]||L||
| |
| − | |-
| |
| − | | PHYE||Histidine kinase||[http://arabidopsis.org/servlets/TairObject?id=26568&type=locus TAIR]||L||[http://www.uniprot.org/uniprot/B9U4G5 Grape]
| |
| − | |-
| |
| − | | PI||Floral homeotic gene encoding a MADS domain transcription factor. Required for the specification of petal and stamen identities.||[http://arabidopsis.org/servlets/TairObject?id=131273&type=locus TAIR]||||[http://www.uniprot.org/uniprot/J9SQF2 Grape]
| |
| − | |-
| |
| − | | PIE1||Encodes a protein similar to ATP-dependent, chromatin-remodeling proteins of the ISWI and SWI2/SNF2 family. Genetic analyses suggest that this gene is involved in multiple flowering pathways. Mutations in PIE1 results in suppression of FLC-mediated delay of flowering and causes early flowering in noninductive photoperiods independently of FLC. PIE1 is required for expression of FLC in the shoot apex but not in the root.Along with ARP6 forms a complex to deposit modified histone H2A.Z at several loci within the genome. This modification alters the expression of the target genes (i.e. FLC, MAF4, MAF6).||[http://arabidopsis.org/servlets/TairObject?id=37964&type=locus TAIR]||V||
| |
| − | |-
| |
| − | | PIF3||Transcription factor interacting with photoreceptors phyA and phyB. Forms a ternary complex in vitro with G-box element of the promoters of LHY, CCA1. Acts as a negative regulator of phyB signalling. It degrades rapidly after irradiation of dark grown seedlings in a process controlled by phytochromes. Does not play a significant role in controlling light input and function of the circadian clockwork. Binds to G- and E-boxes, but not to other ACEs. Binds to anthocyanin biosynthetic genes in a light- and HY5-independent fashion. PIF3 function as a transcriptional activator can be functionally and mechanistically separated from its role in repression of PhyB mediated processes.||[http://arabidopsis.org/servlets/TairObject?id=27554&type=locus TAIR]||||
| |
| − | |-
| |
| − | | PRR7||PRR7 and PRR9 are partially redundant essential components of a temperature-sensitive circadian system. CCA1 and LHY had a positive effect on PRR7 expression levels. Acts as transcriptional repressor of CCA1 and LHY.||[http://arabidopsis.org/servlets/TairObject?id=131538&type=locus TAIR]||L||
| |
| − | |-
| |
| − | | RAV1||Encodes an AP2/B3 domain transcription factor which is upregulated in response to low temperature. It contains a B3 DNA binding domain. It has circadian regulation and may function as a negative growth regulator.||[http://arabidopsis.org/servlets/TairObject?id=137721&type=locus TAIR]||L||
| |
| − | |-
| |
| − | | REF6||Relative of Early Flowering 6 (REF6) encodes a Jumonji N/C and zinc finger domain-containing protein that acts as a positive regulator of flowering in an FLC-dependent pathway. REF6 mutants have hyperacetylation of histone H4 at the FLC locus. REF6 interacts with BES1 in a Y2H assay and in vitro. REF6 may play a role in brassinoteroid signaling by affecting histone methylation in the promoters of BR-responsive genes. It is most closely related to the JHDM3 subfamily of JmjN/C proteins.||[http://arabidopsis.org/servlets/TairObject?id=500230554&type=locus TAIR]||||
| |
| − | |-
| |
| − | | RFI2||Zinc ion binding||[http://arabidopsis.org/servlets/TairObject?id=32070&type=locus TAIR]||L||
| |
| − | |-
| |
| − | | SEC*||Has O-linked N-acetyl glucosamine transferase activity. Similar to Arabidopsis SPY gene.||[http://arabidopsis.org/servlets/TairObject?id=40600&type=locus TAIR]||||
| |
| − | |-
| |
| − | | SEP1||Encodes a stress enhanced protein that localizes to the thylakoid membrane and whose mRNA is upregulated in response to high light intensity. It may be involved in chlorophyll binding.||[http://arabidopsis.org/servlets/TairObject?id=127910&type=locus TAIR]||||
| |
| − | |-
| |
| − | | SEP2||Stress enhanced protein 2 (SEP2) chlorophyll a/b-binding protein||[http://arabidopsis.org/servlets/TairObject?id=33372&type=locus TAIR]||||
| |
| − | |-
| |
| − | | SEP3||Member of the MADs box transcription factor family. SEP3 is redundant with SEP1 and 2. Flowers of SEP1/2/3 triple mutants show a conversion of petals and stamens to sepals.SEP3 forms heterotetrameric complexes with other MADS box family members and binds to the CArG box motif.||[http://arabidopsis.org/servlets/TairObject?id=30252&type=locus TAIR]||||
| |
| − | |-
| |
| − | | SEP4||This gene belongs to the family of SEP genes. It is involved in the development of sepals, petals, stamens and carpels. Additionally, it plays a central role in the determination of flower meristem and organ identity.||[http://arabidopsis.org/servlets/TairObject?id=32207&type=locus TAIR]||||
| |
| − | |-
| |
| − | | SHP1*||One of two genes (SHP1 and SHP2) that are required for fruit dehiscence. The two genes control dehiscence zone differentiation and promote the lignification of adjacent cells.||[http://arabidopsis.org/servlets/TairObject?id=39912&type=locus TAIR]||||
| |
| − | |-
| |
| − | | SHP2*||AGAMOUS [AG]-like MADS box protein (AGL5) involved in fruit development (valve margin and dehiscence zone differentiation). A putative direct target of AG. SHP2 has been shown to be a downstream gene of the complex formed by AG and SEP proteins (SEP4 alone does not form a functional complex with AG).||[http://arabidopsis.org/servlets/TairObject?id=33362&type=locus TAIR]||||
| |
| − | |-
| |
| − | | SMZ||Encodes a AP2 domain transcription factor that can repress flowering. SMZ and its paralogous gene, SNARCHZAPFEN (SNZ), share a signature with partial complementarity to the miR172 microRNA, whose precursor is induced upon flowering.||[http://arabidopsis.org/servlets/TairObject?id=37065&type=locus TAIR]||||
| |
| − | |-
| |
| − | | SNZ||Encodes a AP2 domain transcription factor that can repress flowering. SNZ and its paralogous gene, SCHLAFMUTZE (SMZ), share a signature with partial complementarity to the miR172 microRNA, whose precursor is induced upon flowering.||[http://arabidopsis.org/servlets/TairObject?id=33943&type=locus TAIR]||||
| |
| − | |-
| |
| − | | SOC1||Transcription activator active in flowering time control. May integrate signals from the photoperiod, vernalization and autonomous floral induction pathways. Can modulate class B and C homeotic genes expression. When associated with AGL24, mediates effect of gibberellins on flowering under short-day conditions, and regulates the expression of LEAFY (LFY), which links floral induction and floral development.||[http://www.uniprot.org/uniprot/O64645 UniProtKB]||FPI||[http://www.uniprot.org/uniprot/D1MDP7 Grape], [http://www.uniprot.org/uniprot/G3GAX1 Strawberry]
| |
| − | |-
| |
| − | | SPA1||Encodes a member of the SPA (suppressor of phyA-105) protein family (SPA1-SPA4). SPA proteins contain an N-terminal serine/threonine kinase-like motif followed by a coiled-coil structure and a C-terminal WD-repeat domain. SPA1 is a PHYA signaling intermediate, putative regulator of PHYA signaling pathway. Light responsive repressor of photomorphogenesis. Involved in regulating circadian rhythms and flowering time in plants. Under constant light, the abundance of SPA1 protein exhibited circadian regulation, whereas under constant darkness, SPA1 protein levels remained unchanged. In addition, the spa1-3 mutation slightly shortened circadian period of CCA1, TOC1/PRR1 and SPA1 transcript accumulation under constant light.||[http://arabidopsis.org/servlets/TairObject?id=31336&type=locus TAIR]||L||
| |
| − | |-
| |
| − | | SPA2||Encodes a member of the SPA (suppressor of phyA-105) protein family (SPA1-SPA4). SPA proteins contain an N-terminal serine/threonine kinase-like motif followed by a coiled-coil structure and a C-terminal WD-repeat domain. SPA proteins function redundantly in suppressing photomorphogenesis in dark- and light-grown seedlings. SPA2 primarily regulates seedling development in darkness and has little function in light-grown seedlings or adult plants.||[http://arabidopsis.org/servlets/TairObject?id=129633&type=locus TAIR]||L||
| |
| − | |-
| |
| − | | SPA3||Encodes a member of the SPA (suppressor of phyA-105) protein family (SPA1-SPA4). SPA proteins contain an N-terminal serine/threonine kinase-like motif followed by a coiled-coil structure and a C-terminal WD-repeat domain. SPA proteins function redundantly in suppressing photomorphogenesis in dark- and light-grown seedlings. SPA3 (and SPA4) predominantly regulates elongation growth in adult plants.||[http://arabidopsis.org/servlets/TairObject?id=500230681&type=locus TAIR]||L||
| |
| − | |-
| |
| − | | SPA4||Encodes a member of the SPA (suppressor of phyA-105) protein family (SPA1-SPA4). SPA proteins contain an N-terminal serine/threonine kinase-like motif followed by a coiled-coil structure and a C-terminal WD-repeat domain. SPA proteins function redundantly in suppressing photomorphogenesis in dark- and light-grown seedlings. SPA4 (and SPA3) predominantly regulates elongation growth in adult plants.||[http://arabidopsis.org/servlets/TairObject?id=31031&type=locus TAIR]||L||
| |
| − | |-
| |
| − | | SPL3||Encodes a member of the SPL (squamosa-promoter binding protein-like)gene family, a novel gene family encoding DNA binding proteins and putative transcription factors. Contains the SBP-box, which encodes the SBP-domain, required and sufficient for interaction with DNA. It binds DNA, may directly regulate AP1, and is involved in regulation of flowering and vegetative phase change. Its temporal expression is regulated by the microRNA miR156. The target site for the microRNA is in the 3'UTR.||[http://arabidopsis.org/servlets/TairObject?id=34194&type=locus TAIR]||MI||
| |
| − | |-
| |
| − | | SPY||Encodes a N-acetyl glucosamine transferase that may glycosylate other molecules involved in GA signaling. Contains a tetratricopeptide repeat region, and a novel carboxy-terminal region. SPY acts as both a repressor of GA responses and as a positive regulation of cytokinin signalling. SPY may be involved in reducing ROS accumulation in response to stress.||[http://arabidopsis.org/servlets/TairObject?id=36698&type=locus TAIR]||||
| |
| − | |-
| |
| − | | STK*||Encodes a MADS box transcription factor expressed in the carpel and ovules. Plays a maternal role in fertilization and seed development.||[http://arabidopsis.org/servlets/TairObject?id=130184&type=locus TAIR]||||
| |
| − | |-
| |
| − | | STM||Class I knotted-like homeodomain protein that is required for shoot apical meristem (SAM) formation during embryogenesis and for SAM function throughout the lifetime of the plant. Functions by preventing incorporation of cells in the meristem center into differentiating organ primordia.||[http://arabidopsis.org/servlets/TairObject?id=29436&type=locus TAIR]||||
| |
| − | |-
| |
| − | | SUF4||Encodes SUF4 (SUPPRESSOR of FRI 4), a putative zinc-finger-containing transcription factor that is required for delayed flowering in winter-annual Arabidopsis. suf4 mutations strongly suppress the late-flowering phenotype of FRI (FRIGIDA) mutants. suf4 mutants also show reduced H3K4 trimethylation at FLC (FLOWERING LOCUS C), a floral inhibitor. SUF4 may act to specifically recruit a putative histone H3 methyltransferase EFS (EARLY FLOWERING IN SHORT DAYS) and the PAF1-like complex to the FLC locus.||[http://arabidopsis.org/servlets/TairObject?id=28096&type=locus TAIR]||V||
| |
| − | |-
| |
| − | | SVP||Encodes a nuclear protein that acts as a floral repressor and that functions within the thermosensory pathway. SVP represses FT expression via direct binding to the vCArG III motif in the FT promoter.||[http://arabidopsis.org/servlets/TairObject?id=31694&type=locus TAIR]||Am, V||[http://www.uniprot.org/uniprot/H9CTT8 Grape]
| |
| − | |-
| |
| − | | SWN||Encodes a polycomb group protein. Forms part of a large protein complex that can include VRN2 (VERNALIZATION 2), VIN3 (VERNALIZATION INSENSITIVE 3) and polycomb group proteins FERTILIZATION INDEPENDENT ENDOSPERM (FIE) and CURLY LEAF (CLF). The complex has a role in establishing FLC (FLOWERING LOCUS C) repression during vernalization. Performs a partially redundant role to MEA in controlling seed initiation by helping to suppress central cell nucleus endosperm proliferation within the FG.||[http://arabidopsis.org/servlets/TairObject?id=129104&type=locus TAIR]||Au, V||
| |
| − | |-
| |
| − | | TEM1||Encodes a member of the RAV transcription factor family that contains AP2 and B3 binding domains. Involved in the regulation of flowering under long days. Loss of function results in early flowering. Overexpression causes late flowering and repression of expression of FT. Novel transcriptional regulator involved in ethylene signaling. Promoter bound by EIN3. EDF1 in turn, binds to promoter elements in ethylene responsive genes.||[http://arabidopsis.org/servlets/TairObject?id=30061&type=locus TAIR]||L||
| |
| − | |-
| |
| − | | TEM2||Rav2 is part of a complex that has been named `regulator of the (H+)-ATPase of the vacuolar and endosomal membranes' (RAVE)||[http://arabidopsis.org/servlets/TairObject?id=27570&type=locus TAIR]||L||
| |
| − | |-
| |
| − | | TFL1||Controls inflorescence meristem identity. Involved in the floral initiation process. Ortholog of the Antirrhinum gene CENTRORADIALIS (CEN). Involved in protein trafficking to the protein storage vacuole.||[http://arabidopsis.org/servlets/TairObject?id=131459&type=locus TAIR]||Am, FPI||[http://www.uniprot.org/uniprot/Q84MI7 Grape], [http://www.uniprot.org/uniprot/G5CJU3 Strawberry]
| |
| − | |-
| |
| − | | TOC1||Pseudo response regulator involved in the generation of circadian rhythms. TOC1 appears to shorten the period of circumnutation speed. TOC1 contributes to the plant fitness (carbon fixation, biomass) by influencing the circadian clock period. PRR3 may increase the stability of TOC1 by preventing interactions between TOC1 and the F-box protein ZTL. Expression of TOC1 is correlated with rhythmic changes in chromatin organization.||[http://arabidopsis.org/servlets/TairObject?id=133196&type=locus TAIR]||L||
| |
| − | |-
| |
| − | | TOE1||Pathogenesis-related transcriptional factor/ERF, DNA-binding||[http://arabidopsis.org/servlets/TairObject?id=34025&type=locus TAIR]||||
| |
| − | |-
| |
| − | | TOE2||Sequence-specific DNA binding transcription factor activity; regulation of transcription, DNA-dependent||[http://arabidopsis.org/servlets/TairObject?id=133319&type=locus TAIR]||||
| |
| − | |-
| |
| − | | TOE3||Pathogenesis-related transcriptional factor/ERF, DNA-binding||[http://arabidopsis.org/servlets/TairObject?id=132134&type=locus TAIR]||||
| |
| − | |-
| |
| − | | TSF||Encodes a floral inducer that is a homolog of FT. Plants overexpressing this gene flower earlier than Col. Loss-of-function mutations flower later in short days. TSF and FT play overlapping roles in the promotion of flowering, with FT playing the dominant role.TSF sequences show extensive variation in different accessions and may contribute to quantitative variation in flowering time in these accessions. TSF has a complex pattern of spatial expression; it is expressed mainly in phloem and expression is regulated by daylength and vernalization.||[http://arabidopsis.org/servlets/TairObject?id=26553&type=locus TAIR]||Am, FPI||
| |
| − | |-
| |
| − | | TT16||Encodes a MADS box protein. Regulates proanthocyanidin biosynthesis in the inner-most cell layer of the seed coat. Also controls cell shape of the inner-most cell layer of the seed coat. Also shown to be necessary for determining the identity of the endothelial layer within the ovule. Paralogous to GOA. Plays a maternal role in fertilization and seed development.||[http://arabidopsis.org/servlets/TairObject?id=133655&type=locus TAIR]||||
| |
| − | |-
| |
| − | | UFO||Required for the proper identity of the floral meristem. Involved in establishing the whorled pattern of floral organs, in the control of specification of the floral meristem, and in the activation of APETALA3 and PISTILLATA. UFO is found at the AP3 promoter in a LFY-dependent manner, suggesting that it works with LFY to regulate AP3 expression. UFO may also promote the ubiquitylation of LFY.||[http://arabidopsis.org/servlets/TairObject?id=28094&type=locus TAIR]||||
| |
| − | |-
| |
| − | | ULP1A*||Encodes a deSUMOylating enzyme. In vitro it has both peptidase activity and isopeptidase activity: it can cleave the C-terminal residues from SUMO to activate it for attachment to a target protein and it can also act on the isopeptide bond between SUMO and another protein. In vitro assays suggest that this enzyme is active against SUMO1 and SUMO2. It has weak activity with SUMO3 and cannot act on SUMO5. The N-terminal regulatory region of this protein is required for full activity.||[http://arabidopsis.org/servlets/TairObject?id=36190&type=locus TAIR]||||
| |
| − | |-
| |
| − | | ULP1B*||Cysteine-type peptidase activity; proteolysis||[http://arabidopsis.org/servlets/TairObject?id=128314&type=locus TAIR]||||
| |
| − | |-
| |
| − | | UVR3*||Required for photorepair of 6-4 photoproducts in Arabidopsis thaliana.||[http://arabidopsis.org/servlets/TairObject?id=38932&type=locus TAIR]||L||
| |
| − | |-
| |
| − | | VEL1||Encodes a protein with similarity to VRN5 and VIN3.Contains both a fibronectin III and PHD finger domain. VEL1 is a part of a polycomb repressive complex (PRC2) that is involved in epigenetic silencing of the FLC flowering locus.||[http://arabidopsis.org/servlets/TairObject?id=128580&type=locus TAIR]||Au, V||
| |
| − | |-
| |
| − | | VEL2||Protein of unknown function||[http://arabidopsis.org/servlets/TairObject?id=32230&type=locus TAIR]||Au||
| |
| − | |-
| |
| − | | VEL3||Protein of unknown function||[http://arabidopsis.org/servlets/TairObject?id=32249&type=locus TAIR]||Au||
| |
| − | |-
| |
| − | | VIN3||Encodes a plant homeodomain protein VERNALIZATION INSENSITIVE 3 (VIN3). In planta VIN3 and VRN2, VERNALIZATION 2, are part of a large protein complex that can include the polycomb group (PcG) proteins FERTILIZATION INDEPENDENT ENDOSPERM (FIE), CURLY LEAF (CLF), and SWINGER (SWN or EZA1). The complex has a role in establishing FLC (FLOWERING LOCUS C) repression during vernalization.||[http://arabidopsis.org/servlets/TairObject?id=134483&type=locus TAIR]||Au, V||
| |
| − | |-
| |
| − | | VIP3||The protein is composed of repeats of WD motif which is involved in protein complex formation. The gene is involved in flower timing and flower development. This gene is predicted to encode a protein with a DWD motif. It can bind to DDB1a in Y2H assays, and DDB1b in co-IP assays, and may be involved in the formation of a CUL4-based E3 ubiquitin ligase. Loss of gene function leads to a redistribution of H3K4me3 and K3K36me2 modifications within genes but not a change in the overall abundance of these modifications within chromatin.||[http://arabidopsis.org/servlets/TairObject?id=127868&type=locus TAIR]||||
| |
| − | |-
| |
| − | | VIP4||Encodes highly hydrophilic protein involved in positively regulating FLC expression. Mutants are early flowering and show a loss of FLC expression in the absence of cold.||[http://arabidopsis.org/servlets/TairObject?id=132654&type=locus TAIR]||||
| |
| − | |-
| |
| − | | VRN1||Required for vernalization. Essential for the complete repression of FLC in vernalized plants. Required for the methylation of histone H3||[http://arabidopsis.org/servlets/TairObject?id=37620&type=locus TAIR]||V||
| |
| − | |-
| |
| − | | VRN2||The VERNALIZATION2 (VRN2) gene mediates vernalization and encodes a nuclear-localized zinc finger protein with similarity to Polycomb group (PcG) proteins of plants and animals. In wild-type Arabidopsis, vernalization results in the stable reduction of the levels of the floral repressor FLC. In vrn2 mutants, FLC expression is downregulated normally in response to vernalization, but instead of remaining low, FLC mRNA levels increase when plants are returned to normal temperatures. VRN2 maintains FLC repression after a cold treatment, serving as a mechanism for the cellular memory of vernalization. Required for complete repression of FLC. Required for the methylation of histone H3||[http://arabidopsis.org/servlets/TairObject?id=228106&type=locus TAIR]||Au, V||
| |
| − | |-
| |
| − | | VRN5||Encodes Vernalization Insensitive 3-like 1 (VIL1). VIL1 is involved in the photoperiod and vernalization of Arabidopsis by regulating expression of the related floral repressors Flowering Locus C (FLC) and Flowering Locus M (FLM). VIL1, along with VIN3 (Vernalization Insensitive 3) is necessary for the chromatin modification to FLC and FLM.||[http://arabidopsis.org/servlets/TairObject?id=39196&type=locus TAIR]||Au, V||
| |
| − | |-
| |
| − | | WUS||Homeobox gene controlling the stem cell pool. Expressed in the stem cell organizing center of meristems. Required to keep the stem cells in an undifferentiated state. Regulation of WUS transcription is a central checkpoint in stem cell control. The size of the WUS expression domain controls the size of the stem cell population through WUS indirectly activating the expression of CLAVATA3 (CLV3) in the stem cells and CLV3 repressing WUS transcription through the CLV1 receptor kinase signaling pathway. Repression of WUS transcription through AGAMOUS (AG) activity controls stem cell activity in the determinate floral meristem. Binds to TAAT element core motif. WUS is also involved in cell differentiation during anther development.||[http://arabidopsis.org/servlets/TairObject?id=34708&type=locus TAIR]||||[http://www.uniprot.org/uniprot/C3UZN8 Grape]
| |
| − | |-
| |
| − | | ZTL||Encodes clock-associated PAS protein ZTL; Also known as FKF1-like protein 2 or ADAGIO1(ADO1). A protein containing a PAS domain ZTL contributes to the plant fitness (carbon fixation, biomass) by influencing the circadian clock period. ZTL is the F-box component of an SCF complex implicated in the degradation of TOC1.||[http://arabidopsis.org/servlets/TairObject?id=134481&type=locus TAIR]||L||
| |
| | |} | | |} |
| | + | |
| | + | |
| | + | ==References== |
| | + | *Amasino RM. Seasonal and developmental timing of flowering. ''Plant J,'' '''61,''' 1001-1013 (2010). |
| | + | *Amasino RM. Vernalization and flowering time. ''Curr Opin Plant Biol,'' '''16,''' 154–158 (2005). |
| | + | *Ballerini ES, Kramer EM. Environmental and molecular analysis of the floral transition in the lower eudicot Aquilegia formosa. ''Evodevo,'' '''2,''' 1-20 (2011). |
| | + | *Bäurle I, Dean C. The timing of developmental transitions in plants. ''Cell,'' '''125,''' 655-664 (2006). |
| | + | *Boss PK, Bastow RM, Mylne JS, Dean C. Multiple pathways in the decision to flower: enabling, promoting, and resetting. ''Plant Cell,'' '''16,''' S18-S31 (2004). |
| | + | *Farrona S, Coupland G, Turck F. The impact of chromatin regulation on the floral transition. ''Semin Cell Dev Biol,'' '''19,''' 560-573 (2008). |
| | + | *Ferrier T, Matus JT, Jin J, Riechmann JL. Arabidopsis paves the way: genomic and network analyses in crops. ''Curr Opin Biotechnol,'' '''22,''' 260-270 (2011). |
| | + | *Fornara F, de Montaigu A, Coupland G. SnapShot: Control of flowering in Arabidopsis. ''Cell,'' '''141,''' 550-550.e2 (2010). |
| | + | *Henderson IR, Dean C. Control of Arabidopsis flowering: the chill before the bloom. ''Development,'' '''131,''' 3829-3838 (2004). |
| | + | *He Y, Amasino RM. Role of chromatin modification in flowering-time control. ''Trends Plant Sci,'' '''10,''' 30-35 (2005). |
| | + | *Higgins JA, Bailey PC, Laurie DA. Comparative genomics of flowering time pathways using Brachypodium distachyon as a model for the temperate grasses. ''PLoS One,'' '''5,''' e10065 (2010). |
| | + | *Huala E, Dickerman AW, Garcia-Hernandez M, Weems D, Reiser L, LaFond F, Hanley D, Kiphart D, Zhuang M, Huang W, Mueller LA, Bhattacharyya D, Bhaya D, Sobral BW, Beavis W, Meinke DW, Town CD, Somerville C, Rhee SY. The Arabidopsis Information Resource (TAIR): a comprehensive database and web-based information retrieval, analysis, and visualization system for a model plant. ''Nucleic Acids Res,'' '''29,''' 102-105 (2001). |
| | + | *Izawa T, Takahashi Y, Yano M. Comparative biology comes into bloom: genomic and genetic comparison of flowering pathways in rice and Arabidopsis. ''Curr Opin Plant Biol,'' '''6,''' 113-120 (2003). |
| | + | *Jung CH, Wong CE, Singh MB, Bhalla PL. Comparative genomic analysis of soybean flowering genes. ''PLoS One,'' '''7,''' e38250 (2012). |
| | + | *Kim DH, Sung S. The Plant Homeo Domain finger protein, VIN3-LIKE 2, is necessary for photoperiod-mediated epigenetic regulation of the floral repressor, MAF5. ''Proc Natl Acad Sci USA,'' '''107,''' 17029-17034 (2010). |
| | + | *Kim SY, Lee J, Eshed-Williams L, Zilberman D, Sung ZR. EMF1 and PRC2 cooperate to repress key regulators of Arabidopsis development. ''PLoS Genet,'' '''8,''' e1002512 (2012). |
| | + | *Liu C, Thong Z, Yu H. Coming into bloom: the specification of floral meristems. ''Development,'' '''136,''' 3379-3391 (2009). |
| | + | *Posé D, Yant L, Schmid M. The end of innocence: flowering networks explode in complexity. ''Curr Opin Plant Biol'' '''15,''' 45-50 (2012). |
| | + | *Schneitz K, Balasubramanian S. Floral Meristems. ''eLS'' (John Wiley & Sons Ltd, Chichester, 2009). |
| | + | *SSR Tool. ''Genome Database for Vaccinium.'' |
| | + | *Sung ZR, Chen L, Moon YH, Lertpiriyapong K. Mechanisms of floral repression in Arabidopsis. ''Curr Opin Plant Biol,'' '''6,''' 29-35 (2003). |
| | + | *Taiz L, Zeiger E. ''Plant Physiology Fifth Edition Ch. 20'' (Sinauer Associates, Sunderland, MA, 2010). |
| | + | *Turck F, Adrian J. A lesson in complexity: regulation of FLOWERING LOCUS T. ''Max Planck Institute for Plant Breeding Research.'' |
| | + | *UniProt Consortium. Reorganizing the protein space at the Universal Protein Resource (UniProt). ''Nucleic Acids Res,'' '''40,''' D71-D75 (2012). |
| | + | *Wellmer F, Riechmann JL. Gene networks controlling the initiation of flower development. ''Trends Genet,'' '''26,''' 519-527 (2010). |
| | + | *Yamaguchi A, Kobayashi Y, Goto K, Abe M, Araki T. TWIN SISTER OF FT (TSF) acts as a floral pathway integrator redundantly with FT. ''Plant Cell Physiol,'' '''46,''' 1175-1189 (2005). |
| | + | *Yu S, Galvão VC, Zhang YC, Horrer D, Zhang TQ, Hao YH, Feng YQ, Wang S, Schmid M, Wang JW. Gibberellin regulates the Arabidopsis floral transition through miR156-targeted SQUAMOSA promoter binding-like transcription factors. ''Plant Cell,'' '''24,''' 3320-3332 (2012). |
| | + | *Zhang JZ, Ai XY, Sun LM, Zhang DL, Guo WW, Deng XX, Hu CG. Transcriptome profile analysis of flowering molecular processes of early flowering trifoliate orange mutant and the wild-type [Poncirus trifoliata (L.) Raf.] by massively parallel signature sequencing. ''BMC Genomics,'' '''12,''' 63 (2011). |
This table lists 108 genes involved in the age, ambient temperature, autonomous, gibberellin, light signaling, polycomb, and vernalization pathways, almost all of which "have been implicated in flowering-time control based on isolation of loss-of-function mutations or analysis of transgenic plants." (Fornara et al., 2010)
| Arabidopsis Locus |
Other Names |
AA Source |
Pathway |
Top Hit Vaccinium Scaffold |
E Value
|
| AT1G01060 |
LATE ELONGATED HYPOCOTYL, LATE ELONGATED HYPOCOTYL 1, LHY, LHY1 |
TAIR |
Light Signaling |
Scaffold00140 (length 354209) at 234299 |
2e-19
|
| AT1G02580 |
EMB173, EMBRYO DEFECTIVE 173, FERTILIZATION INDEPENDENT SEED 1, FIS1, MEA, MEDEA, SDG5, SET DOMAIN-CONTAINING PROTEIN 5 |
TAIR |
Polycomb |
Scaffold00354 (length 215005) at 64805 |
4e-17
|
| AT1G04400 |
AT-PHH1, ATCRY2, CRY2, CRYPTOCHROME 2, FHA, PHH1 |
TAIR |
Light Signaling |
Scaffold00649 (length 159319) at 28296 |
1e-119
|
| AT1G09570 |
ELONGATED HYPOCOTYL 8, FAR RED ELONGATED 1, FAR RED ELONGATED HYPOCOTYL 2, FHY2, FRE1, HY8, PHYA, PHYTOCHROME A |
TAIR |
Light Signaling |
Scaffold03861 (length 7403) at 3771 |
0.0
|
| AT1G13260 |
EDF4, ETHYLENE RESPONSE DNA BINDING FACTOR 4, RAV1, RELATED TO ABI3/VP1 1 |
TAIR |
Light Signaling |
Scaffold00930 (length 110378) at 63069 |
1e-103
|
| AT1G14920 |
GAI, GIBBERELLIC ACID INSENSITIVE, RESTORATION ON GROWTH ON AMMONIA 2, RGA2 |
TAIR |
Gibberellin |
Scaffold01360 (length 81306) at 51382 |
0.0
|
| AT1G20330 |
COTYLEDON VASCULAR PATTERN 1, CVP1, FRILL1, FRL1, SMT2, STEROL METHYLTRANSFERASE 2 |
TAIR |
Vernalization |
Scaffold02142 (length 46662) at 24743 |
9e-06
|
| AT1G22770 |
FB, GI, GIGANTEA |
TAIR |
Light Signaling |
Scaffold00100 (length 346620) at 198329 |
0.0
|
| AT1G25560 |
EDF1, ETHYLENE RESPONSE DNA BINDING FACTOR 1, TEM1, TEMPRANILLO 1 |
TAIR |
Light Signaling |
Scaffold00930 (length 110378) at 63096 |
3e-99
|
| AT1G26790 |
|
TAIR |
Light Signaling |
Scaffold00079 (length 471015) at 69048 |
6e-33
|
| AT1G27370 |
SPL10, SQUAMOSA PROMOTER BINDING PROTEIN-LIKE 10 |
TAIR |
Age |
Scaffold00127 (length 336847) at 165806 |
5e-33
|
| AT1G29160 |
|
TAIR |
Light Signaling |
Scaffold00270 (length 246371) at 215229 |
4e-50
|
| AT1G30970 |
SUF4, SUPPRESSOR OF FRIGIDA4 |
TAIR |
Vernalization |
Scaffold00348 (length 210978) at 75279 |
1e-15
|
| AT1G31814 |
FRIGIDA LIKE 2, FRL2 |
TAIR |
Vernalization |
Scaffold00289 (length 259591) at 150896 |
2e-19
|
| AT1G47250 |
20S PROTEASOME ALPHA SUBUNIT F2, PAF2 |
TAIR |
Vernalization |
Scaffold00528 (length 197283) at 110933 |
3e-64
|
| AT1G53090 |
SPA1-RELATED 4, SPA4 |
TAIR |
Light Signaling |
Scaffold00734 (length 158513) at 143450 |
3e-129
|
| AT1G53160 |
FLORAL TRANSITION AT THE MERISTEM6, FTM6, SPL4, SQUAMOSA PROMOTER BINDING PROTEIN-LIKE 4 |
TAIR |
Age |
Scaffold00062 (length 451336) at 412489 |
1e-15
|
| AT1G65480 |
FLOWERING LOCUS T, FT |
TAIR |
Ambient Temperature |
Scaffold00357 (length 260075) at 58774 |
6e-35
|
| AT1G66350 |
RGA-LIKE 1, RGL, RGL1 |
TAIR |
Gibberellin |
Scaffold00134 (length 346346) at 174707 |
0.0
|
| AT1G68050 |
"FLAVIN-BINDING, KELCH REPEAT, F BOX 1", ADO3, FKF1 |
TAIR |
Light Signaling |
Scaffold00110 (length 358202) at 120279 |
0.0
|
| AT1G68840 |
ATRAV2, EDF2, ETHYLENE RESPONSE DNA BINDING FACTOR 2, RAP2.8, RAV2, RELATED TO ABI3/VP1 2, RELATED TO AP2 8, TEM2, TEMPRANILLO 2 |
TAIR |
Light Signaling |
Scaffold00930 (length 110378) at 63096 |
5e-102
|
| AT1G77080 |
AGAMOUS-LIKE 27, AGL27, FLM, FLOWERING LOCUS M, MADS AFFECTING FLOWERING 1, MAF1 |
TAIR |
Ambient Temperature, Vernalization |
Scaffold10765 (length 2447) at 744 |
3e-13
|
| AT1G77300 |
ASH1 HOMOLOG 2, ASHH2, CAROTENOID CHLOROPLAST REGULATORY1, CCR1, EARLY FLOWERING IN SHORT DAYS, EFS, LAZ2, LAZARUS 2, SDG8, SET DOMAIN GROUP 8 |
TAIR |
Vernalization |
Scaffold00894 (length 114877) at 89213 |
1e-27
|
| AT2G01570 |
REPRESSOR OF GA, REPRESSOR OF GA1-3 1, RGA, RGA1 |
TAIR |
Gibberellin |
Scaffold01360 (length 81306) at 51382 |
0.0
|
| AT2G06255 |
ELF4-L3, ELF4-LIKE 3 |
TAIR |
Light Signaling |
Scaffold00336 (length 230445) at 188137 |
1e-39
|
| AT2G17770 |
ATBZIP27, BASIC REGION/LEUCINE ZIPPER MOTIF 27, BZIP27, FD PARALOG, FDP |
TAIR |
Ambient Temperature |
Scaffold00367 (length 240396) at 113139 |
7e-17
|
| AT2G18790 |
HY3, OOP1, OUT OF PHASE 1, PHYB, PHYTOCHROME B |
TAIR |
Light Signaling |
Scaffold00751 (length 152548) at 83698 |
0.0
|
| AT2G18870 |
VEL3, VERNALIZATION5/VIN3-LIKE 3, VIL4, VIN3-LIKE 4 |
TAIR |
Autonomous |
Scaffold00396 (length 187979) at 73810 |
7e-19
|
| AT2G18880 |
VEL2, VERNALIZATION5/VIN3-LIKE 2, VIL3, VIN3-LIKE 3 |
TAIR |
Autonomous |
Scaffold00026 (length 499904) at 369396 |
6e-41
|
| AT2G18915 |
ADAGIO 2, ADO2, LKP2, LOV KELCH PROTEIN 2 |
TAIR |
Light Signaling |
Scaffold00026 (length 499904) at 445582 |
0.0
|
| AT2G19520 |
ACG1, ATMSI4, FVE, MSI4, MULTICOPY SUPPRESSOR OF IRA1 4, NFC04, NFC4 |
TAIR |
Ambient Temperature, Autonomous |
Scaffold00728 (length 157708) at 8716 |
6e-21
|
| AT2G22540 |
AGAMOUS-LIKE 22, AGL22, SHORT VEGETATIVE PHASE, SVP |
TAIR |
Ambient Temperature, Vernalization |
Scaffold01187 (length 101153) at 47701 |
2e-24
|
| AT2G23380 |
CLF, CURLY LEAF, ICU1, INCURVATA 1, SDG1, SET1, SETDOMAIN 1, SETDOMAIN GROUP 1 |
TAIR |
Autonomous, Polycomb, Vernalization |
Scaffold00354 (length 215005) at 64799 |
5e-41
|
| AT2G25930 |
EARLY FLOWERING 3, ELF3, PYK20 |
TAIR |
Light Signaling |
Scaffold00371 (length 223748) at 73111 |
6e-19
|
| AT2G32950 |
ARABIDOPSIS THALIANA CONSTITUTIVE PHOTOMORPHOGENIC 1, ATCOP1, CONSTITUTIVE PHOTOMORPHOGENIC 1, COP1, DEETIOLATED MUTANT 340, DET340, EMB168, EMBRYO DEFECTIVE 168, FUS1, FUSCA 1 |
TAIR |
Light Signaling |
Scaffold00734 (length 158513) at 142779 |
5e-62
|
| AT2G33810 |
SPL3, SQUAMOSA PROMOTER BINDING PROTEIN-LIKE 3 |
TAIR |
Age |
Scaffold00062 (length 451336) at 412489 |
2e-17
|
| AT2G33835 |
FES1, FRIGIDA-ESSENTIAL 1 |
TAIR |
Vernalization |
Scaffold00102 (length 383343) at 88264 |
7e-17
|
| AT2G34140 |
|
TAIR |
Light Signaling |
Scaffold00270 (length 246371) at 215217 |
7e-49
|
| AT2G35670 |
FERTILIZATION INDEPENDENT SEED 2, FERTILIZATION-INDEPENDENT ENDOSPERM 2, FIE2, FIS2 |
TAIR |
Polycomb |
Scaffold00857 (length 126644) at 55565 |
3e-04
|
| AT2G40080 |
EARLY FLOWERING 4, ELF4 |
TAIR |
Light Signaling |
Scaffold00254 (length 265810) at 228805 |
4e-24
|
| AT2G42200 |
ATSPL9, SPL9, SQUAMOSA PROMOTER BINDING PROTEIN-LIKE 9 |
TAIR |
Age |
Scaffold00691 (length 141413) at 61795 |
5e-21
|
| AT2G43410 |
FPA |
TAIR |
Autonomous |
Scaffold01689 (length 75472) at 47663 |
4e-45
|
| AT2G46340 |
SPA1, SUPPRESSOR OF PHYA-105 1 |
TAIR |
Light Signaling |
Scaffold00734 (length 158513) at 142779 |
3e-69
|
| AT2G46790 |
APRR9, ARABIDOPSIS PSEUDO-RESPONSE REGULATOR 9, PRR9, PSEUDO-RESPONSE REGULATOR 9, TL1, TOC1-LIKE PROTEIN 1 |
TAIR |
Light Signaling |
Scaffold00001 (length 1030549) at 322801 |
1e-36
|
| AT2G46830 |
ATCCA1, CCA1, CIRCADIAN CLOCK ASSOCIATED 1 |
TAIR |
Light Signaling |
Scaffold00140 (length 354209) at 234299 |
7e-20
|
| AT2G47700 |
RED AND FAR-RED INSENSITIVE 2, RFI2 |
TAIR |
Light Signaling |
Scaffold01059 (length 101143) at 96557 |
1e-19
|
| AT3G02380 |
ATCOL2, B-BOX DOMAIN PROTEIN 3, BBX3, COL2, CONSTANS-LIKE 2 |
TAIR |
Light Signaling |
Scaffold01843 (length 51900) at 40767 |
3e-48
|
| AT3G03450 |
RGA-LIKE 2, RGL2 |
TAIR |
Gibberellin |
Scaffold00134 (length 346346) at 174722 |
0.0
|
| AT3G04610 |
FLK, FLOWERING LOCUS KH DOMAIN |
TAIR |
Autonomous |
Scaffold01384 (length 81068) at 28309 |
4e-29
|
| AT3G05120 |
ATGID1A, GA INSENSITIVE DWARF1A, GID1A |
TAIR |
Gibberellin |
Scaffold00101 (length 425332) at 398762 |
4e-175
|
| AT3G07650 |
B-BOX DOMAIN PROTEIN 7, BBX7, COL9, CONSTANS-LIKE 9 |
TAIR |
Light Signaling |
Scaffold00832 (length 123094) at 89599 |
1e-55
|
| AT3G10390 |
FLD, FLOWERING LOCUS D |
TAIR |
Ambient Temperature, Autonomous |
Scaffold00232 (length 253418) at 139565 |
0.0
|
| AT3G12810 |
CHR13, PHOTOPERIOD-INDEPENDENT EARLY FLOWERING 1, PIE1, SRCAP |
TAIR |
Vernalization |
Scaffold00147 (length 345970) at 30164 |
0.0
|
| AT3G15270 |
SPL5, SQUAMOSA PROMOTER BINDING PROTEIN-LIKE 5 |
TAIR |
Age |
Scaffold00105 (length 392037) at 147284 |
5e-16
|
| AT3G15354 |
SPA1-RELATED 3, SPA3 |
TAIR |
Light Signaling |
Scaffold00734 (length 158513) at 143450 |
1e-144
|
| AT3G18990 |
REDUCED VERNALIZATION RESPONSE 1, REM39, REPRODUCTIVE MERISTEM 39, VRN1 |
TAIR |
Vernalization |
Scaffold00811 (length 108354) at 96661 |
2e-25
|
| AT3G20740 |
FERTILIZATION-INDEPENDENT ENDOSPERM, FERTILIZATION-INDEPENDENT ENDOSPERM 1, FIE, FIE1, FIS3 |
TAIR |
Autonomous, Polycomb, Vernalization |
Scaffold01670 (length 60304) at 9836 |
5e-20
|
| AT3G21320 |
|
TAIR |
Light Signaling |
Scaffold00509 (length 192554) at 181887 |
1e-21
|
| AT3G24440 |
VERNALIZATION 5, VIL1, VIN3-LIKE 1, VRN5 |
TAIR |
Autonomous, Vernalization |
Scaffold00396 (length 187979) at 73795 |
2e-69
|
| AT3G25730 |
EDF3, ETHYLENE RESPONSE DNA BINDING FACTOR 3 |
TAIR |
Light Signaling |
Scaffold00930 (length 110378) at 63060 |
5e-100
|
| AT3G33520 |
ACTIN-RELATED PROTEIN 6, ARP6, ATARP6, EARLY IN SHORT DAYS 1, ESD1, SUF3, SUPPRESSOR OF FRI 3 |
TAIR |
Ambient Temperature, Vernalization |
Scaffold00246 (length 286612) at 268521 |
1e-53
|
| AT3G46640 |
LUX, LUX ARRHYTHMO, PCL1, PHYTOCLOCK 1 |
TAIR |
Light Signaling |
Scaffold01150 (length 88293) at 80272 |
3e-34
|
| AT3G47500 |
CDF3, CYCLING DOF FACTOR 3 |
TAIR |
Light Signaling |
Scaffold00079 (length 471015) at 68175 |
1e-85
|
| AT4G00650 |
FLA, FLOWERING LOCUS A, FRI, FRIGIDA |
TAIR |
Vernalization |
Scaffold00039 (length 505275) at 216632 |
4e-50
|
| AT4G02020 |
EZA1, SDG10, SET DOMAIN-CONTAINING PROTEIN 10, SWINGER, SWN |
TAIR |
Autonomous, Polycomb, Vernalization |
Scaffold00354 (length 215005) at 64799 |
2e-48
|
| AT4G02560 |
LD, LUMINIDEPENDENS |
TAIR |
Autonomous |
Scaffold00002 (length 840149) at 158601 |
4e-28
|
| AT4G08920 |
ATCRY1, BLU1, BLUE LIGHT UNINHIBITED 1, CRY1, CRYPTOCHROME 1, ELONGATED HYPOCOTYL 4, HY4, OOP2, OUT OF PHASE 2 |
TAIR |
Light Signaling |
Scaffold00331 (length 261439) at 124817 |
4e-128
|
| AT4G11110 |
SPA1-RELATED 2, SPA2 |
TAIR |
Light Signaling |
Scaffold01034 (length 107107) at 19205 |
2e-62
|
| AT4G11880 |
AGAMOUS-LIKE 14, AGL14 |
TAIR |
Vernalization |
Scaffold00249 (length 258199) at 131012 |
2e-29
|
| AT4G16250 |
PHYD, PHYTOCHROME D |
TAIR |
Light Signaling |
Scaffold00751 (length 152548) at 83707 |
0.0
|
| AT4G16280 |
FCA |
TAIR |
Ambient Temperature, Autonomous |
Scaffold00104 (length 360147) at 120318 |
5e-13
|
| AT4G16845 |
REDUCED VERNALIZATION RESPONSE 2, VRN2 |
TAIR |
Autonomous, Polycomb, Vernalization |
Scaffold00857 (length 126644) at 55203 |
3e-07
|
| AT4G18130 |
PHYE, PHYTOCHROME E |
TAIR |
Light Signaling |
Scaffold00751 (length 152548) at 83749 |
0.0
|
| AT4G20370 |
TSF, TWIN SISTER OF FT |
TAIR |
Ambient Temperature |
Scaffold00357 (length 260075) at 58807 |
1e-33
|
| AT4G22950 |
AGAMOUS-LIKE 19, AGL19, GL19 |
TAIR |
Vernalization |
Scaffold00249 (length 258199) at 131012 |
4e-28
|
| AT4G24540 |
AGAMOUS-LIKE 24, AGL24 |
TAIR |
Vernalization |
Scaffold01187 (length 101153) at 47701 |
2e-25
|
| AT4G26000 |
PEP, PEPPER |
TAIR |
Vernalization |
Scaffold00021 (length 527983) at 157278 |
3e-29
|
| AT4G30200 |
VEL1, VERNALIZATION5/VIN3-LIKE 1, VIL2, VIN3-LIKE 2 |
TAIR |
Autonomous, Vernalization |
Scaffold00026 (length 499904) at 369417 |
7e-64
|
| AT4G34530 |
CIB1, CRYPTOCHROME-INTERACTING BASIC-HELIX-LOOP-HELIX 1 |
TAIR |
Light Signaling |
Scaffold01322 (length 99420) at 82955 |
2e-29
|
| AT4G35900 |
ATBZIP14, FD, FD-1 |
TAIR |
Ambient Temperature |
Scaffold00367 (length 240396) at 113139 |
1e-16
|
| AT5G02810 |
APRR7, PRR7, PSEUDO-RESPONSE REGULATOR 7 |
TAIR |
Light Signaling |
Scaffold00125 (length 356885) at 276518 |
1e-33
|
| AT5G03840 |
TERMINAL FLOWER 1, TFL1 |
TAIR |
Ambient Temperature |
Scaffold00181 (length 337602) at 12825 |
1e-28
|
| AT5G08330 |
CCA1 HIKING EXPEDITION, CHE, TRANSCRIPTION FACTOR TCP21, TCP21 |
UniProtKB |
Light Signaling |
Scaffold00993 (length 109486) at 294 |
5e-27
|
| AT5G10140 |
AGAMOUS-LIKE 25, AGL25, FLC, FLF, FLOWERING LOCUS C, FLOWERING LOCUS F |
TAIR |
Ambient Temperature, Vernalization |
Scaffold10765 (length 2447) at 708 |
8e-22
|
| AT5G11530 |
EMBRYONIC FLOWER 1, EMF1 |
TAIR |
Polycomb |
Scaffold00253 (length 240879) at 11069 |
1e-06
|
| AT5G13480 |
FY |
TAIR |
Autonomous |
Scaffold00166 (length 328538) at 74796 |
2e-43
|
| AT5G15840 |
B-BOX DOMAIN PROTEIN 1, BBX1, CO, CONSTANS, FG |
TAIR |
Light Signaling |
Scaffold01843 (length 51900) at 40767 |
3e-29
|
| AT5G15850 |
ATCOL1, B-BOX DOMAIN PROTEIN 2, BBX2, COL1, CONSTANS-LIKE 1 |
TAIR |
Light Signaling |
Scaffold01843 (length 51900) at 40767 |
5e-44
|
| AT5G17690 |
ATLHP1, LHP1, LIKE HETEROCHROMATIN PROTEIN 1, TERMINAL FLOWER 2, TFL2 |
TAIR |
Polycomb |
Scaffold00696 (length 140617) at 128983 |
7e-13
|
| AT5G23150 |
ENHANCER OF AG-4 2, HUA2 |
TAIR |
Vernalization |
Scaffold00686 (length 145647) at 102689 |
5e-42
|
| AT5G24470 |
APRR5, PRR5, PSEUDO-RESPONSE REGULATOR 5 |
TAIR |
Light Signaling |
Scaffold00001 (length 1030549) at 322798 |
3e-40
|
| AT5G24930 |
ATCOL4, B-BOX DOMAIN PROTEIN 5, BBX5, COL4, CONSTANS-LIKE 4 |
TAIR |
Light Signaling |
Scaffold11225 (length 3326) at 2816 |
8e-23
|
| AT5G35840 |
PHYC, PHYTOCHROME C |
TAIR |
Light Signaling |
Scaffold01070 (length 102706) at 75395 |
0.0
|
| AT5G37055 |
ATSWC6, SEF, SERRATED LEAVES AND EARLY FLOWERING |
TAIR |
Vernalization |
Scaffold00925 (length 141061) at 38143 |
4e-29
|
| AT5G39660 |
CDF2, CYCLING DOF FACTOR 2 |
TAIR |
Light Signaling |
Scaffold00651 (length 145047) at 19066 |
1e-90
|
| AT5G42790 |
ARS5, ARSENIC TOLERANCE 5, ATPSM30, PAF1, PROTEASOME ALPHA SUBUNIT F1 |
TAIR |
Vernalization |
Scaffold00528 (length 197283) at 110933 |
5e-64
|
| AT5G51230 |
ATEMF2, CYR1, CYTOKININ RESISTANT 1, EMBRYONIC FLOWER 2, EMF2, VEF2 |
TAIR |
Polycomb |
Scaffold00857 (length 126644) at 44887 |
9e-19
|
| AT5G57360 |
ADAGIO 1, ADO1, FKF1-LIKE PROTEIN 2, FKL2, LKP1, LOV KELCH PROTEIN 1, ZEITLUPE, ZTL |
TAIR |
Light Signaling |
Scaffold00026 (length 499904) at 445597 |
0.0
|
| AT5G57380 |
VERNALIZATION INSENSITIVE 3, VIN3 |
TAIR |
Autonomous, Vernalization |
Scaffold00026 (length 499904) at 369405 |
3e-48
|
| AT5G57660 |
ATCOL5, B-BOX DOMAIN PROTEIN 6, BBX6, COL5, CONSTANS-LIKE 5 |
TAIR |
Light Signaling |
Scaffold11225 (length 3326) at 2816 |
1e-25
|
| AT5G58230 |
ARABIDOPSIS MULTICOPY SUPRESSOR OF IRA1, ATMSI1, MATERNAL EFFECT EMBRYO ARREST 70, MEE70, MSI1, MULTICOPY SUPRESSOR OF IRA1 |
TAIR |
Autonomous, Polycomb, Vernalization |
Scaffold00615 (length 179658) at 161081 |
2e-61
|
| AT5G60100 |
APRR3, PRR3, PSEUDO-RESPONSE REGULATOR 3 |
TAIR |
Light Signaling |
Scaffold02075 (length 43325) at 14139 |
3e-25
|
| AT5G61380 |
APRR1, ATTOC1, PRR1, PSEUDO-RESPONSE REGULATOR 1, TIMING OF CAB EXPRESSION 1, TOC1 |
TAIR |
Light Signaling |
Scaffold00753 (length 123742) at 70468 |
2e-47
|
| AT5G62430 |
CDF1, CYCLING DOF FACTOR 1 |
TAIR |
Light Signaling |
Scaffold01102 (length 99830) at 51435 |
1e-59
|
| AT5G65050 |
AGAMOUS-LIKE 31, AGL31, MADS AFFECTING FLOWERING 2, MAF2 |
TAIR |
Ambient Temperature, Vernalization |
Scaffold10765 (length 2447) at 723 |
4e-20
|
| AT5G65060 |
AGAMOUS-LIKE 70, AGL70, FCL3, MADS AFFECTING FLOWERING 3, MAF3 |
TAIR |
Ambient Temperature, Vernalization |
Scaffold10765 (length 2447) at 639 |
7e-15
|
| AT5G65070 |
AGAMOUS-LIKE 69, AGL69, FCL4, MADS AFFECTING FLOWERING 4, MAF4 |
TAIR |
Ambient Temperature, Vernalization |
Scaffold10765 (length 2447) at 723 |
6e-20
|
| AT5G65080 |
AGAMOUS-LIKE 68, AGL68, MADS AFFECTING FLOWERING 5, MAF5 |
TAIR |
Ambient Temperature, Vernalization |
Scaffold10765 (length 2447) at 699 |
1e-18
|