Blasting Spacers for Our Genome
Blasting Spacers
EXPLORATION OF THE IMPORTANT FINDINGS
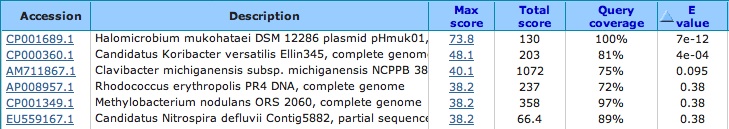
CRISPR one results from CRISPR finder are shown below. I blasted all of the spacers using the nr/nt database and the results are shown below.
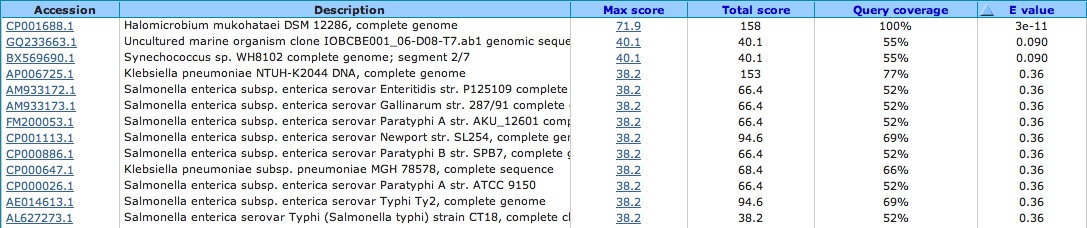
Spacer One: TGCGTCGTCCGGTGGCCGTCAATAAATGTCGCAAGGG
No significant hits . . . lowest E-value is 5.9
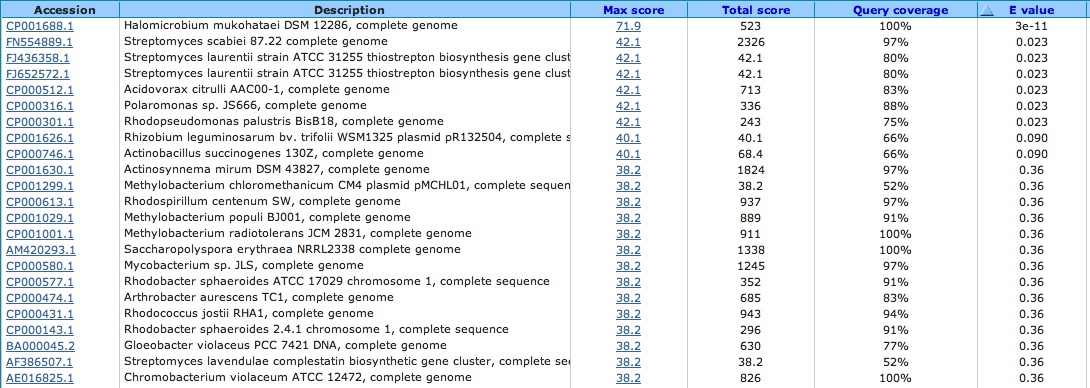
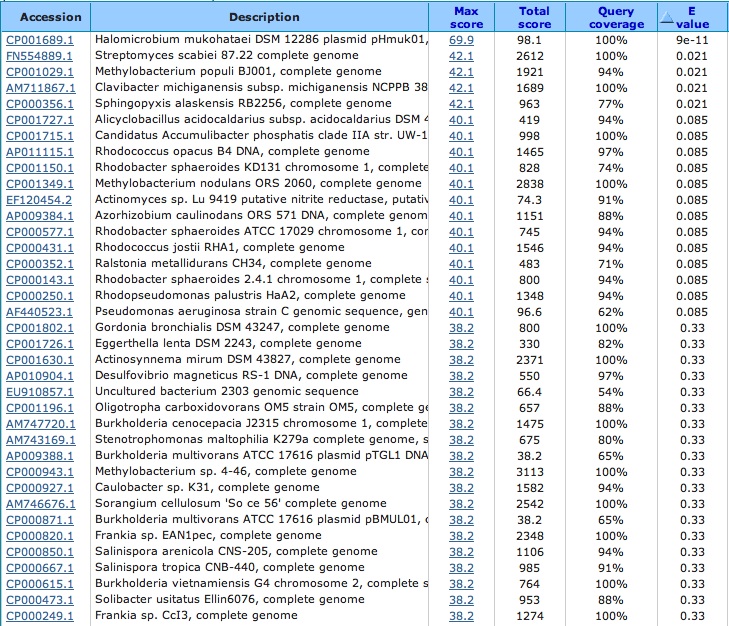
Spacer Two: TCCTACGACCTCGTCGGCGTCAACGGCTGGCCCGA
Archaeal BJ1 virus complete genome: Good Article
This is the protein sequence location that this match comes from in the virus:
Blasted this little sequence alignment section. . . did not get any significant viral alignments except its own Did blastp to determine if hypothetical protein in archaeal BJ1 virus had hits in other viruses:
The rest of the hits are bacteria and are in coding protein sequences such as ABC transporter and ATP-binding protein.
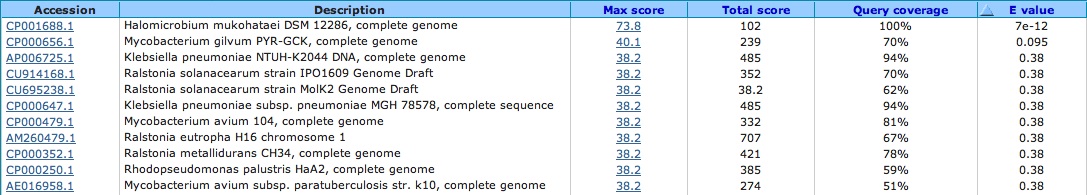
Spacer Three: CACCCTACAACAGGTGAAATCTACCAGACAAAAGA
All of these hits are bacteria and the bacteria hits are part of coding proteins such as conserved hypothetical protein; putative membrane protein, which I did a blastp for and found no viral matches
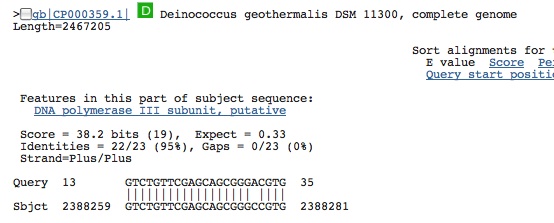
Spacer Four: TCACCCAAGCGCAAGCAACAGCTGATCGAGGACCTG
Hits come from prokaryotes and is from a coding segment
Spacer Five: GCGACGGCGGCCAGTTCCGCGAGGGCGGGAAGGTCC
The following hits are the only significant hits that come from a prokaryote but are n a coding segment of DNA
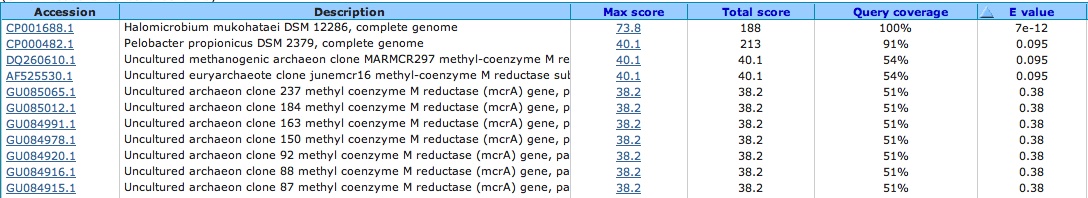
Spacer Six: TGCGAGTGTTGCGGGGAACCGACTCGGGTAGGCCAG
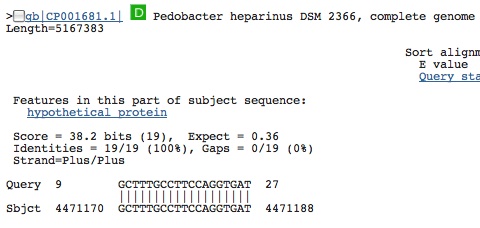
Aligns with something in our genome. . . NOT another spacer but a segment of DNA that flanks two genes and is non-coding. . .
Spacer Seven: ACGGTTTCGCTACCACCATCGCCACCAGCAACTGCCG
Hit from a prokaryote that is in its coding region
Spacer Eight: GACGAGTATGCCAACCGGCTCGTGAGCGGGCGC
No significant hits . . . lowest E-value is 1.2
Spacer Nine: TGGTAGGCGTCGTAGGTGTTCGTGGCGAGCGTGTC
Hit from prokaryote that is in its coding region
Spacer Ten: ACTCACTGGATTATGACCCCTACAACGAGGGCGTCA
No significant hits . . . lowest E-value is 1.4
Spacer Eleven: GGATCTCGATCGTTGTAGTATCCATAGCTGCTATACC
Hits from gene segment in prokaryote or promoter sequence because flanking two genes
Spacer Twelve: GAAGTAACGCAACTCCAGTGAGCGCTACTGAGAGCCC
No significant hits . . . lowest E-value is 1.5
Spacer Thirteen:CCGATCACGCCCTGCCGATACTGGTAGTTCGCGATA
Hit from prokaryote in gene segment
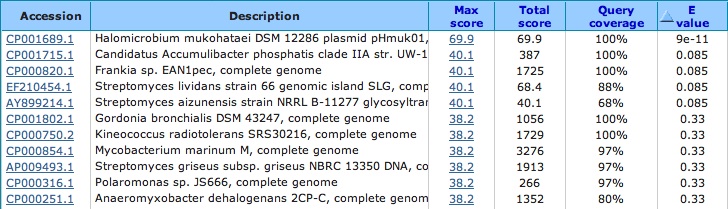
Spacer Fourteen: TCGTCGGCCGGCTCGTCGGCCGACGTGGACTTGC
first hit from prokaryote gene region 2nd hit: from virus
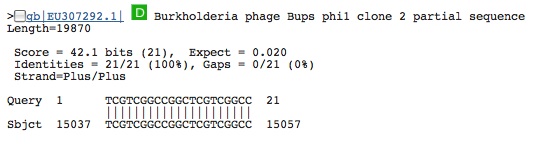
this virus when you blast it. . . it infects pathogens, bacteria and plant. . .maybe archaea too????
Spacer Fifteen: AGTAGGTCTAATGTCTCTCTGTCGTCTATCAGCCCCG
No significant hits . . . lowest E-value is 23
Spacer Sixteen: GCTCTCCGGTGTCACAGGTCAGGTCACGGTCTCCGC
No significant hits . . . lowest E-value is 5.6
Spacer Seventeen: ACGGACAAGTCATCCACCCGCCAGTATCTCCCGGT
Bad hit from prokaryote coding sequence
Spacer Eighteen: AAATACGATCCTGCGGTGACGCTACGTCCGGGGCAGC
No significant hits . . . lowest E-value is 1.5
Spacer Nineteen: CAGATGTGGGGTCTGTGGCCACAGTCTAACATCTCT
No significant hits . . . lowest E-value is 1.4
Spacer Twenty: CCTGATAACGGACTCTTGTAGGTCCGTTAGGTCGT
No significant hits . . . lowest E-value is 5.2
Spacer Twenty-One: AAAAATGAGTGACGTAGACATTCGGCAAAATGCCGG
No significant hits . . . lowest E-value is 1.4
Spacer Twenty-Two: CAGCAGCGAAACGAGCCGTCCGTCCTTTTGAGACA
Hit from prokaryote in gene segment
Spacer Twenty-Three: GTCTAGCCCAGTCTGGTCGGGGTGGTCGGCAGGATCGG
Bad -values= throw out
Spacer Twenty-Four: CACTCCTCATATGTCTGTTCGAGCAGCGGGACGTG
Hit from prokaryote in gene segment
Spacer Twenty-Five: ACCGTTGCCGCCGATCGGCAGCGAGCCGGTGATGTGT
Hits from prokaryotes in flanking sequence or in gene sequence. . . check to see if this is in spacer!!
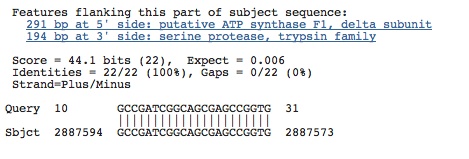
Spacer Twenty-Six: TCAAAGCGAGCCTCGAACGCGACGACGAAGATATG
Hits from prokaryotes in gene segment
Spacer Twenty-Seven: TCCTCCTTGTACCCACGGTCTTGCCGATCCATCCCG
No significant hits . . . lowest E-value is 1.4
Spacer Twenty-Eight: GACTGGCGTGTTGCCGTTCAGGCCGGCGTTGATCCCG
Hits from prokaryotes in gene segment
Spacer Twenty-Nine: CTCAGCAGCAGTCAACGGCATTTTATACACCTTGT
No significant hits . . . lowest E-value is 1.3
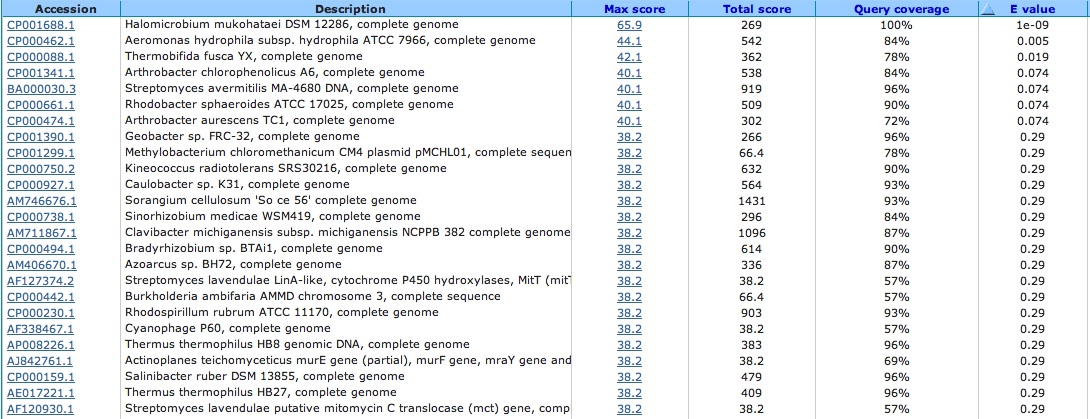
Spacer Thirty: CACCCCTTCCGGGGAGACGAGGAAACCCCGGACGA
Bad e-value and top hit is a bunch of gene from bacteria
Spacer Thirty-One: GTCACGCTGTCTGACGATATGGCTGACCAGGTGC
Hit from prokaryotes in gene segment
Spacer Thirty-Two: CACTCCTGGGCGGCCTCATCGGCGGCCATCGTC
Hit from prokaryotes in gene segment
Spacer Thirty-Three: GTTGTGTGAGGTATGCGATGGACACCACCGATCACG
No significant hits . . . lowest E-value is 1.4
CRISPR Two Spacers: CHANGED parameters to be 1.-1 match/mistatch to make it less intense. . . also excluded eukaryotes NOW
Spacer One: TCCGAGACGTGTTCCCTCTCTAGCTGTGCATCTTCC
Hits from prokaryote in gene segment
Spacer Two: CAGATCTAAAACAATGTCATACGGAAAAATCGACATC
No significant hits . . . lowest E-value is 1.5
Spacer Three: GATCCGGAATATGAAGTGACGAACGATCCGGATACGG
E-values not good and prokaryotes
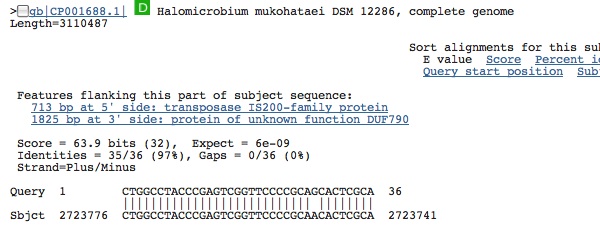
Spacer Four: TCGACGAGATCGGCGCGAACTCGTTCGCTGATACT
Hits from prokaryote in gene segment
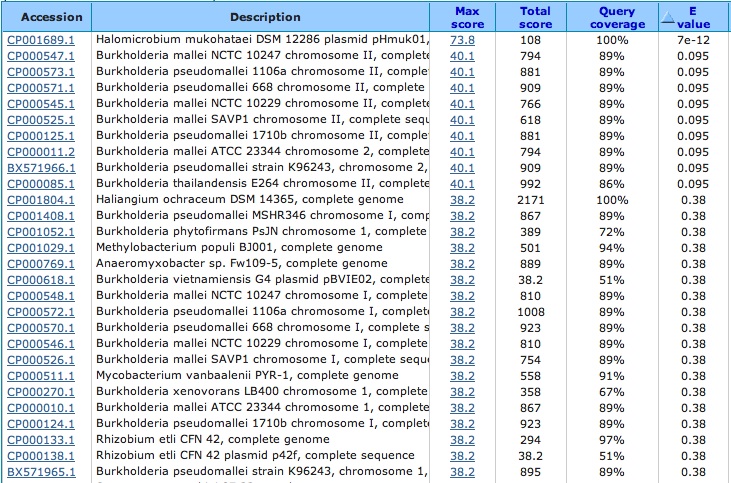
Spacer Five: TCGGGGACCGAGACGACGGGGCCGGGTGCTGTCT
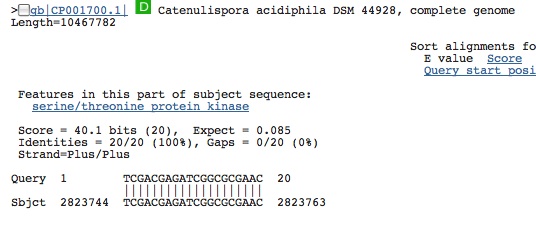
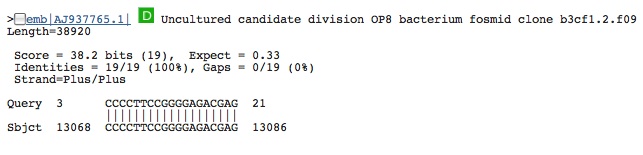
Hit from a virus (same virus even though called different names):
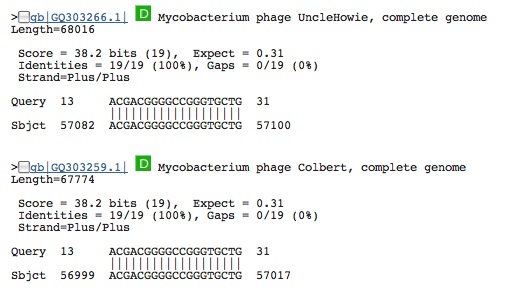
Spacer Six: CCGGAGGGGCCGCTGCGTGGGTGATCTGGAGAGAAGA
E-values not good and from prokaryotes
Spacer Seven: GTTGCGTGAGCTAGCGAAACACCGAGTCCGTGTGAT
No significant hits . . . lowest E-value is 5.5
Spacer Eight: ACGGAAATCCAGCCGATCACCCTCCGAGAGGAGAGG
Hit from prokaryote in gene segment
Spacer Nine: CGGACATTCAGAAGCGCCTGACTAACCGCATGGCT
No significant hits . . . lowest E-value is 5.2
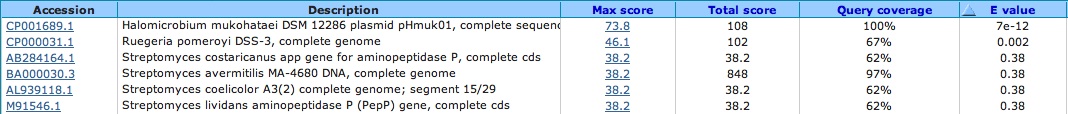
Spacer Ten: CGGGAAGACCACGACCGCCCGCGCCCTCCAGTTCGA
First hit from bacteria in gene segment. . . Second hit is from another archaea in a conserved hypothetical protein. . . is this a true protein. . .??
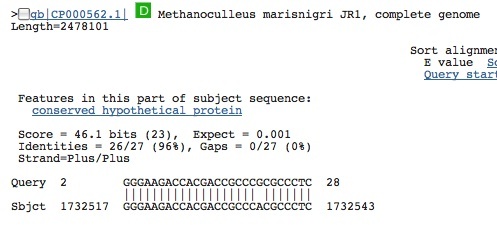
Spacer Eleven: GGCTTCTACGTCGGCAACCGGACCGAGGACGGCGATG
Hit with significant e-value is from a bacteria in a gene segment
Spacer Twelve: AAGTACGCCTCGATCATCAACGGCGTCCGGGCTGT
E-values no good and from a prokaryote in a gene
Spacer Thirteen: GATGCTGTTGAGCTCGTAGCGATCCCACTCGGCGT
No significant hits . . . lowest E-value is 1.3
Spacer Fourteen: AGCCCTTGTGCAATGATCGGGAGTGCAATCCGACC
No significant hits . . . lowest E-value is 5.2
Spacer Fifteen: TCAGGCGAGTTGTCGGACGAACAGCTTGAAGCGTGT
No significant hits . . . lowest E-value is 1.4
Spacer Sixteen: TCGTCGAGCGGCAGGCCGCCGACCGCTACGGGCTG
Hits from prokaryotes in gene segment
Spacer Seventeen: CGACACGGTCGAAGATGCTGGCAGCTCGCGAATCGC
Bad e-value
Spacer Eighteen: GGATCTCGACGGCCGCGTGGCCGTGCATCTCGGGGTC
Hit from prokaryote in gene segment
Spacer Nineteen: AACGCTTCAACGCGCTCTATTGACCGAGCGTATCG
No significant hits . . . lowest E-value is 1.3
Spacer Twenty: AGATCTCGCGGATAAGCTGCCCCCGCCCTCCCATGAG
No significant hits . . . lowest E-value is 5.9
Spacer Twenty-One: CACTTCGAGCGGACTTTGGGCCACCCCGGAAAGTCAG
No significant hits . . . lowest E-value is 1.5
Spacer Twenty-Two: CGATACGTCCGGGACGCCCGTGACGACCACCACTGC
Bad e-value
Spacer Twenty-Three: GGAGGAGCGGATGGACATGAGCGACACGACGATCCG
No significant hits . . . lowest E-value is 1.4
Spacer Twenty-Four: GAGCATCTCTCCAATCAGCGGTCATCAACCGCGA
Hits from prokaryotes in gene segment
Spacer Twenty-Five: AGTATCTGTCTACGCGATACCGCACCGTCAGAGTG
No significant hits . . . lowest E-value is 1.3
Spacer Twenty-Six: TACGACCTTCCCACTGAGGGTCTTGAGCTAACGATT
No significant hits . . . lowest E-value is 5.6
Spacer Twenty-Seven: GCGACGTGCTTACCTCTGACGCCAATATTGACCTT
No significant hits . . . lowest E-value is 5.2
Spacer Twenty-Eight: TTCATCAAGAGGCACACAAGCATGGTGCGTCCAAA
No significant hits . . . lowest E-value is 5.2
Spacer Twenty-Nine: CTGGCCTACCCGAGTCGGTTCCCCGCAGCACTCGCA
Spacer Thirty: ACGATCGTCACCGACACCCTCGGGGCCGGCGGCGC
Hits from prokaryotes in gene segment
Spacer Thirty-One: GAAACGACCGAGACGCAAAGCGAGTTCACACAACTC
No significant hits . . . lowest E-value is 5.6
Spacer Thirty-Two: TTAGATGATCAGGTAGCCTGCTACCAGTGCAGCTGC
Poor e-values
Spacer Thirty-Three: GAGACCCAGCTTTGCCTTCCAGGTGATCAGCTCGTA
Hits from prokaryotes in gene segment
Spacer Thirty-Four: TCGAAGCGCTCGGTCGCGACGGAGACCAGCGACCAGCTG
weird because two hits from our species and these 2 hits are in a conserved protein sequence and not the spacer??????????
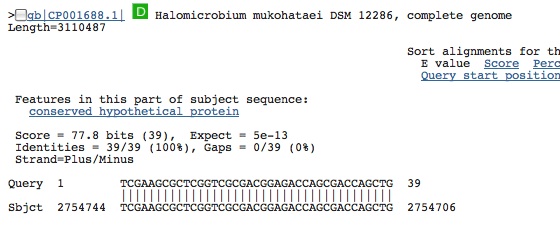
Spacer Thirty-Five: TGACGACCACACACACGAGGCCGTGCGTGTGCTTGTA
Hits from prokaryotes in gene segment
Spacer Thirty-Six: CCACGTCCCGGTGACACGCAGCTCGGTGAGATCGC
Hits from prokaryotes in gene segment
Spacer Thirty-Seven: CATGGAGTCTTCAACATTTCATGGGCTGGGCTTGGCC
No significant hits . . . lowest E-value is 5.9
Spacer Thirty-Eight: CGCAACCCGACGATCGAGGACGGGCCGTCCCTGGA
Bad e-values
Spacer Thirty-Nine: GCGTCGGACTGCGTCGATAGTGTTCGTGCTCATGTT
No significant hits . . . lowest E-value is 1.4
Spacer Forty: CAGACTTCTACTGGAAGGCGAAAACTGAGAAGGCA
No significant hits . . . lowest E-value is 5.2
Spacer Forty-One: TACGCTCGACGACCTCCGTCGTGCGCTCCAGAAGTCA
hit from prokaryote in gene segment
Spacer Forty-Two: GCGCTGCGGACGTGGTGTCAGAGGGGTTACCAGTAACT
No significant hits . . . lowest E-value is 1.6
Spacer Forty-Three: ACTCCGGGTACACTGGTGGCGATGCTCTACTCGCC
No significant hits . . . lowest E-value is 1.3
Spacer Forty-Four: ACACGTTTCTTTTTTTCAGGAGCCATCACTCACTC
No significant hits . . . lowest E-value is 1.3
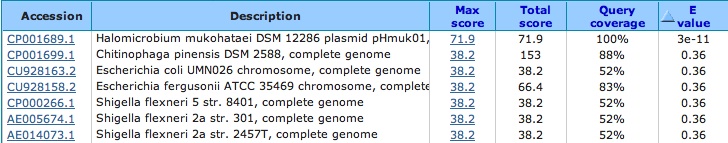
Other things i want to do:
compare our species spacers with themselves look to see if these blast results show hits to any other halophiles look at spacer length and number between our species and others?? any known viruses that infect halophiles??
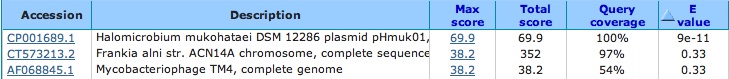
blast the known viral genome against the nr/nt database and it comes up with matches: