Difference between revisions of "Can we solve a 3-SAT problem with supressor logic?"
(→What is suppressor logic?) |
(→What is suppressor logic?) |
||
| Line 16: | Line 16: | ||
A frameshift is a genetic mutation caused by the addition or deletion of nucelotides to a given sequence which codes for a protein. Since codons are read in a series of three, the addition or deletion of nucleotides will disrupt the reading frame. This disruption will most likely cause the production of a nonfunctional protein. | A frameshift is a genetic mutation caused by the addition or deletion of nucelotides to a given sequence which codes for a protein. Since codons are read in a series of three, the addition or deletion of nucleotides will disrupt the reading frame. This disruption will most likely cause the production of a nonfunctional protein. | ||
| − | <center> [[Image: | + | <center> [[Image:suppressor4.GIF]]<br></center> |
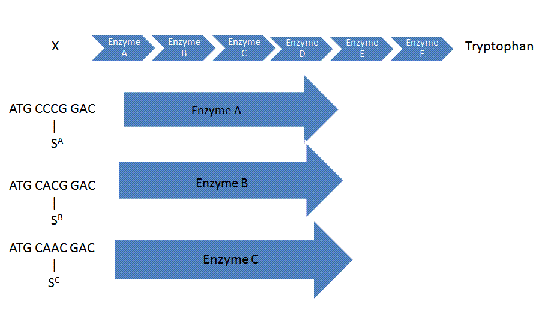
A frameshift occurs and, in this case, a guanine is added to the sequence. If nothing is done, enzyme A will not be made, meaning the clause will not be satisfied. | A frameshift occurs and, in this case, a guanine is added to the sequence. If nothing is done, enzyme A will not be made, meaning the clause will not be satisfied. | ||
Revision as of 20:31, 7 April 2009
Contents
What is the 3-SAT problem?
How did Sakamoto et al. use a DNA computer to solve a 3-SAT problem?
What is suppressor logic?
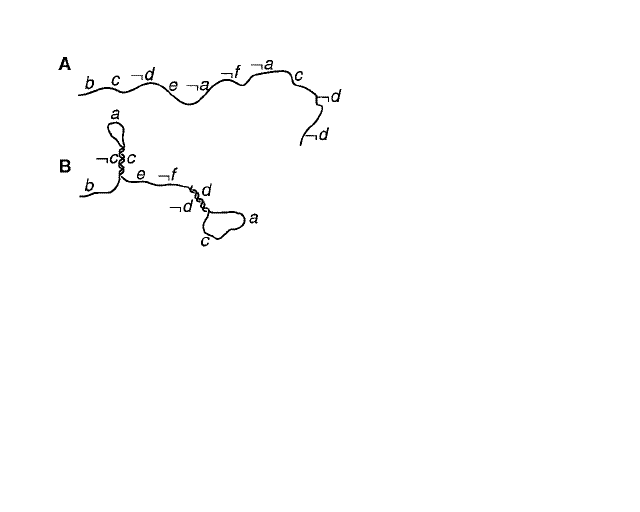
Suppressor Suppressor logic uses suppressor tRNAs to avoid frameshift mutations in an amino acid sequence.
A frameshift is a genetic mutation caused by the addition or deletion of nucelotides to a given sequence which codes for a protein. Since codons are read in a series of three, the addition or deletion of nucleotides will disrupt the reading frame. This disruption will most likely cause the production of a nonfunctional protein.

A frameshift occurs and, in this case, a guanine is added to the sequence. If nothing is done, enzyme A will not be made, meaning the clause will not be satisfied.
The suppressor tRNA allows the 4 letter sequence to be read as a single codon, therefore, keeping the protein on track.
How could suppressor logic be used to solve the Sakamoto 3-SAT problem?
Definitions
Inputs = framshift suppressor tRNAs
Input value = supp a is 1, supp g is 0; supp b is 1, supp h is 0, etc. up to tth 6th pair of f and l
Logical clause (LC) = three inputs connected by OR, eg. (a OR b OR e)
Logical expression (LE) = string of LCs connected by AND
Subroutine
1. Individual bacteral cells use Hin/hix system to randomly choose of of the 64 possible combinations of 6 inputs.
2. Each bacterial cell carries out the following subroutine on each LC: IF LC=TRUE THEN "check the next LC" ELSEIF LC=FALSE "go get a new set of inputs with step 1"
3. If/when a bacterial cell finds a set of inputs that satisfies the entire LE (ie. a solution to the 3-SAT problem), it will glow green.