Difference between revisions of "Explaining My Project"
From GcatWiki
(→Counting Kmers to Tell you about Genome) |
(→Counting Kmers to Tell you about Genome) |
||
| Line 10: | Line 10: | ||
*Genome Coverage = use gamma fit on the good multiplicity values of the best kmer (usually largest). The peak of this line gives genome coverage (see red line) (here about 47.11x) | *Genome Coverage = use gamma fit on the good multiplicity values of the best kmer (usually largest). The peak of this line gives genome coverage (see red line) (here about 47.11x) | ||
*Genome size = number of unique good multiplicity kmers/coverage | *Genome size = number of unique good multiplicity kmers/coverage | ||
| + | *1st peak (@ low multiplicity) = from seq errors | ||
| + | *2nd peak = multiple copies of the same location in the genome | ||
| + | **If a k-mer occurs n times in the genome, we would expect to see it n times as often in the sequencing, so there should be additional peaks for k-mers that occur in repeats. | ||
Latest revision as of 20:30, 8 June 2011
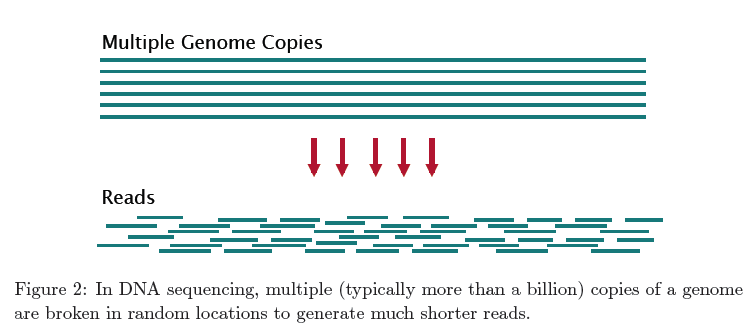
Shotgun Sequencing
Counting Kmers to Tell you about Genome
(taken from https://banana-slug.soe.ucsc.edu/bioinformatic_tools:jellyfish)
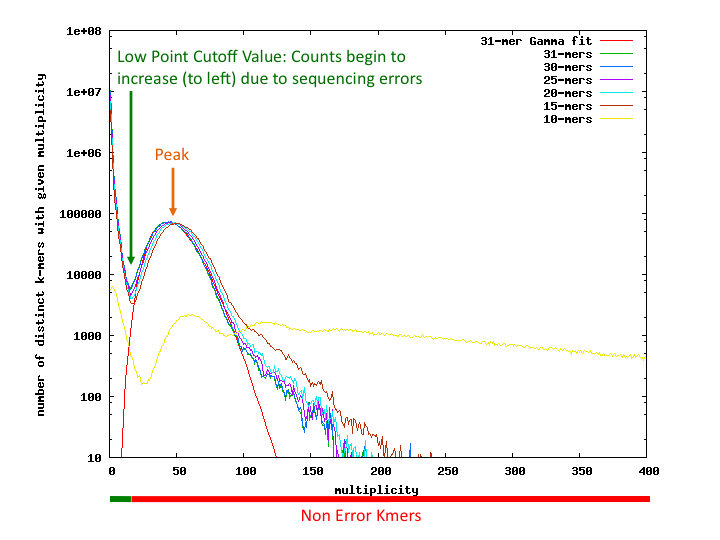
- Bad kmer rate = bad multiplicity kmers/total number of all kmers
- Seq Error Rate = bad kmer Rate/kmer size
- Genome Coverage = use gamma fit on the good multiplicity values of the best kmer (usually largest). The peak of this line gives genome coverage (see red line) (here about 47.11x)
- Genome size = number of unique good multiplicity kmers/coverage
- 1st peak (@ low multiplicity) = from seq errors
- 2nd peak = multiple copies of the same location in the genome
- If a k-mer occurs n times in the genome, we would expect to see it n times as often in the sequencing, so there should be additional peaks for k-mers that occur in repeats.