How to deal with multi-named genes
One always must be leery of the possibility that a gene may have multiple names. When investigating a gene, information can be overlooked by assuming a gene only has one name across species. This tutorial demonstrates how to determine if a gene is referred to by multiple names across species.
Contents
Finding Your Gene of Interest
Say you just are sitting in Biology class and your teacher mentions ATP synthase has a subunit within its complex that is resistant to oligomyocin called Oligomyocin Sensitivity Conferring Protein, gene name OSCP. This sparks your interest, and you want to investigate this gene more. After class you immediately run home, and go to NCBI Gene and search for "OSCP."
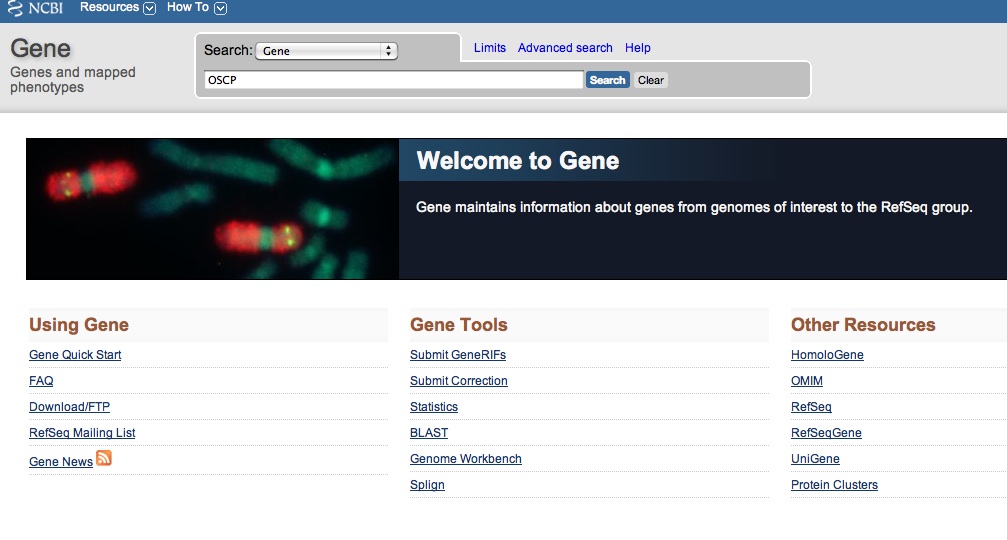
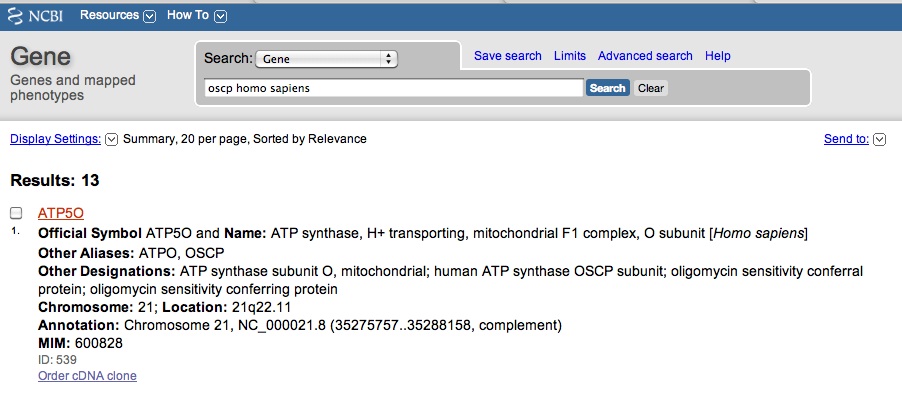
Perfect! You get a hit for OSCP in Drosophila melanogaster. You review the page and realize you want to research OSCP in other organisms, including humans. You then NCBI Gene search "OSCP homo sapiens." You get hits, but none specifically say OSCP. The first hit is for a gene called ATP50.
Is It the Same Gene?
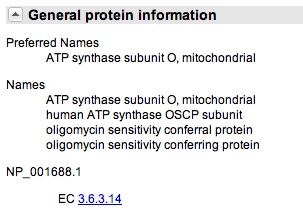
You click onto the ATP50 link and you scroll down to "General Protein Information," as see that gene does encode for oligomyocin sensitivity conferring protein. This suggest that ATP50 is the OSCP in humans, just referred to by another name.
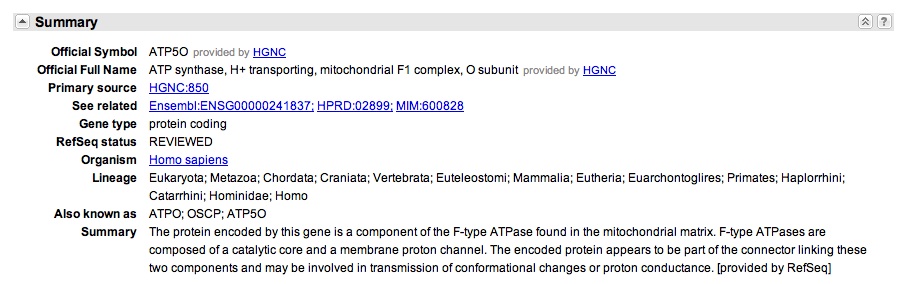
You also notice that there "Also Known As" section, that indicates the gene goes by the names: ATPO; OSCP; ATP5O.

More Databases for More Verification
GeneCards
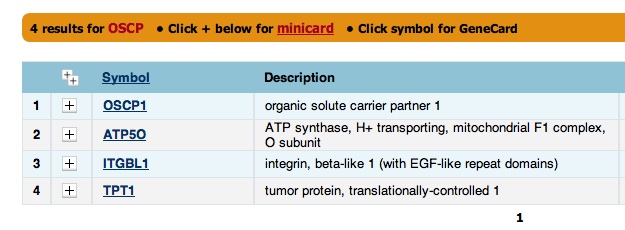
The presence of OSCP in the "Also Known As" section is a little strange because the Drosophila OSCP page did not include ATP50 or ATP0 in the "Also Known As" section. You decide to get further confirmation that OSCP is ATP50. You search "OSCP" on GeneCards, a database for human genes, and get "ATP50" as a hit. That seems promising.
BLAST
You decide you want to find OSCP in other organisms, but do not want to go through the hassel of searching all of NCBI for alternative names, like you did for Homo sapiens. From NCBI Gene, you again go to the human ATP5O page and find the FASTA nucleotide sequence located under "Genomic regions, transcripts, and products". You can either copy and paste the entire sequence into the BLAST box or use the Ref number. The Ref number in this case is NC_000021.8 for ATP50. This time, you choose to copy and paste the nucleotide sequence.

Then go to BLAST and paste the FASTA sequence into the BLAST box. Click BLAST and you compare the nucleotide sequence of ATP5O against all organisms in the BLAST database. The name of the protein the nucleotide sequence hit encodes is clearly shown, making it easy to detect whether or not you have discovered more ATP5O/OSCP genes that encode OSCP protein.
Summary
Gene may have one or more names across species. As in the case with OSCP on the Drosophila page, the alternative name for the gene are not alway listed. Utilize as many database such a NCBI, Pubmed, GeneCard and Blast to discover if there is variance in the names.