(a.k.a. The Lancelator)
Sometimes biologists want to synthesize DNA sequences that are too short for PCR. Thus, they send them off to a large-scale gene assembly company and pay them a minimum order price of approximately $500. Alternatively, they could order single-stranded oligonucleotides of their sequence from a DNA synthesis company and anneal them themselves. This practice eliminates the gene assembly middleman and saves them money since the price of the oligos is based on the number of base pairs. But how does one ensure successful annealing?
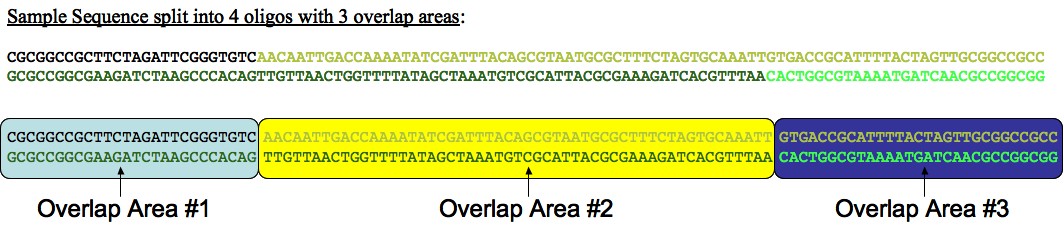
We define an overlap area to exist between breaks on either strand or between an end of the sequence and a break on either strand. As you can see from the figure below, a sequence assembled from 4 oligos will have 3 overlap areas. By maximizing the melting temperatures of our overlap areas across-the-board, we can improve our chances for successful annealing.

The melting temperature for each overlap area is determined by the number of base pairs, or length, and the percentage of G's and C's in the section. As the length of the sequence or the G/C content increases, so does the melting temperature. So in choosing the locations for our breaks, we must balance the length of an overlap area with its G/C content in order to ensure that all of our areas have similar melting temperatures. This website determines the optimal gene assembly for a sequence by calculating the maximin (maximum of the minimums) of the melting temperatures for all possible combinations of overlap areas. It accepts user inputs for the minimum and maximum lengths of overlap areas. The program will not allow you to use overlap lengths less than 20 base pairs or greater than 80. If you want to use lengths that are outside this range, please download the power-user MATLAB or Perl code.
Enter one strand of your sequence in the box. All non-DNA characters A, C, G, and T will be stripped before computing cut.
Web page developed by Lance Harden (laharden[at]davidson[dot]edu), Davidson College '09,
during summer research with Dr. A. Malcolm Campbell (macampbell[at]davidson[dot]edu) and Dr.
Laurie J. Heyer (laheyer[at]davidson[dot]edu) in conjunction with iGEM 2006.
Last update:
July 10, 2008